Translating Genomics to the Clinic
(selected publications)
2024
 (opens in new tab)
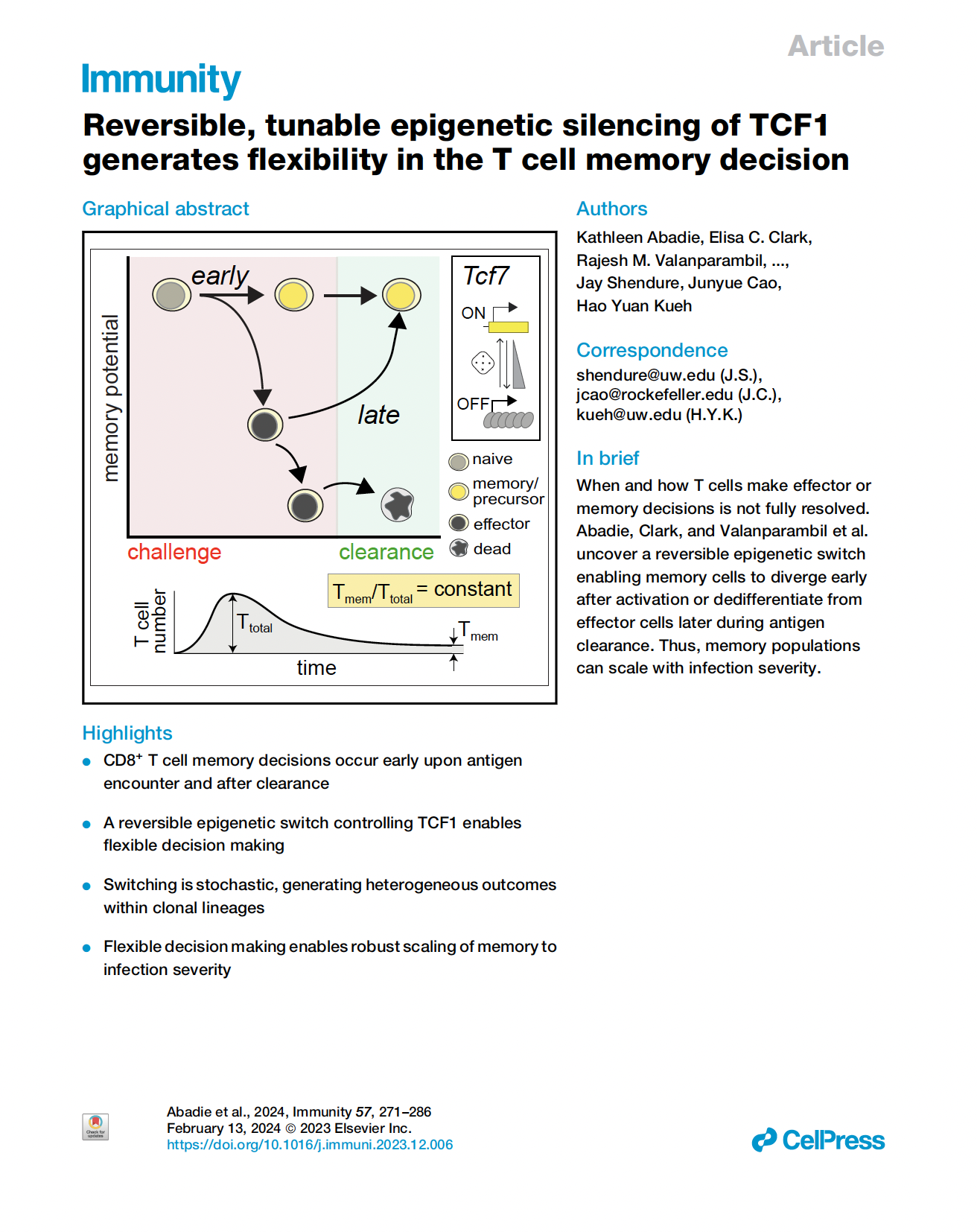
(opens in new tab) Abadie, Daza et al. Reversible, tunable epigenetic silencing of TCF-1 generates flexibility in the T cell memory decision.
Cell Press (2024)
2022
 (opens in new tab)
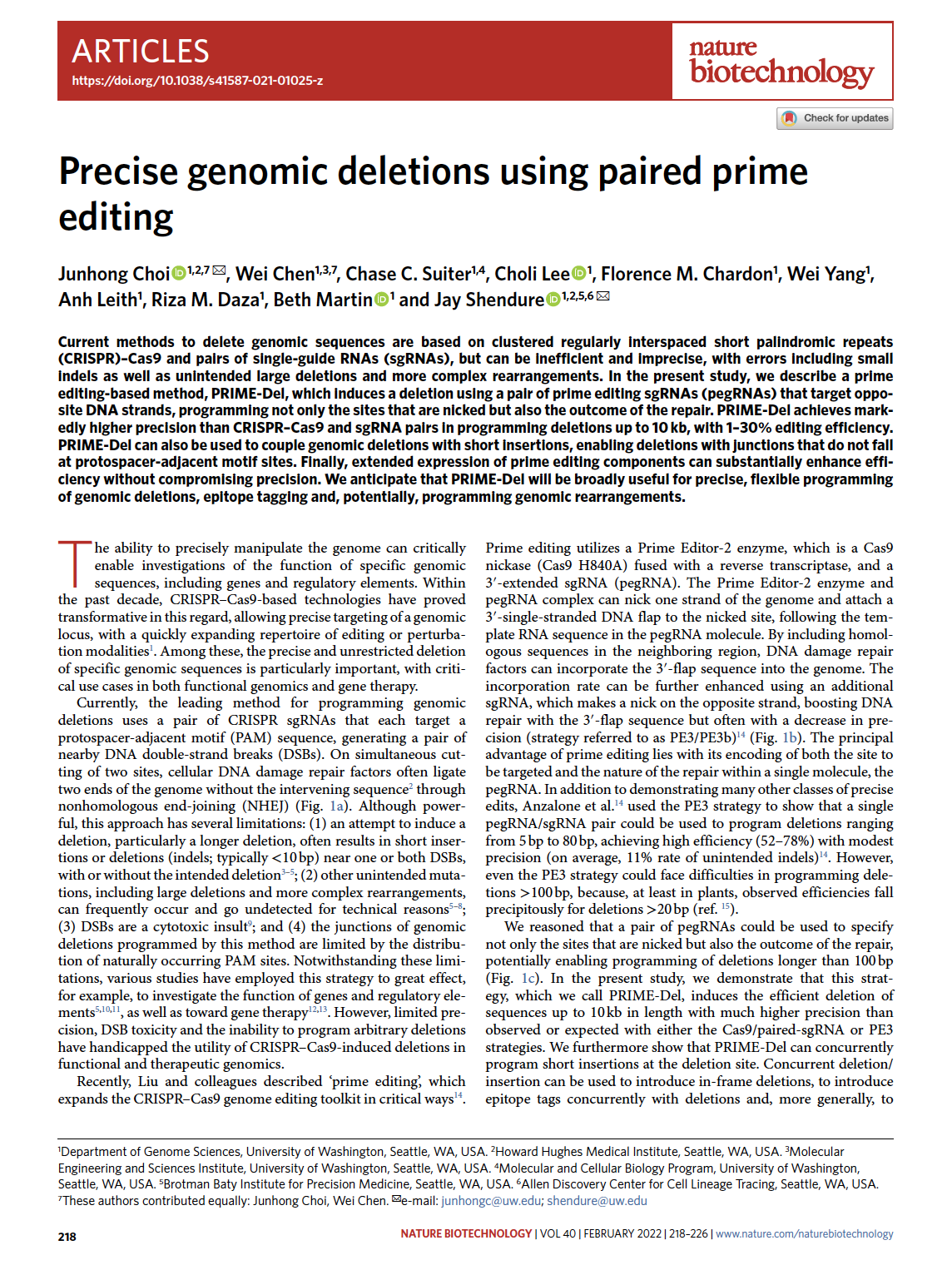
(opens in new tab) Choi, Chen et al. Precise genomic deletions using paired prime editing.
Nature Biotechnology (2022)
2021
 (opens in new tab)
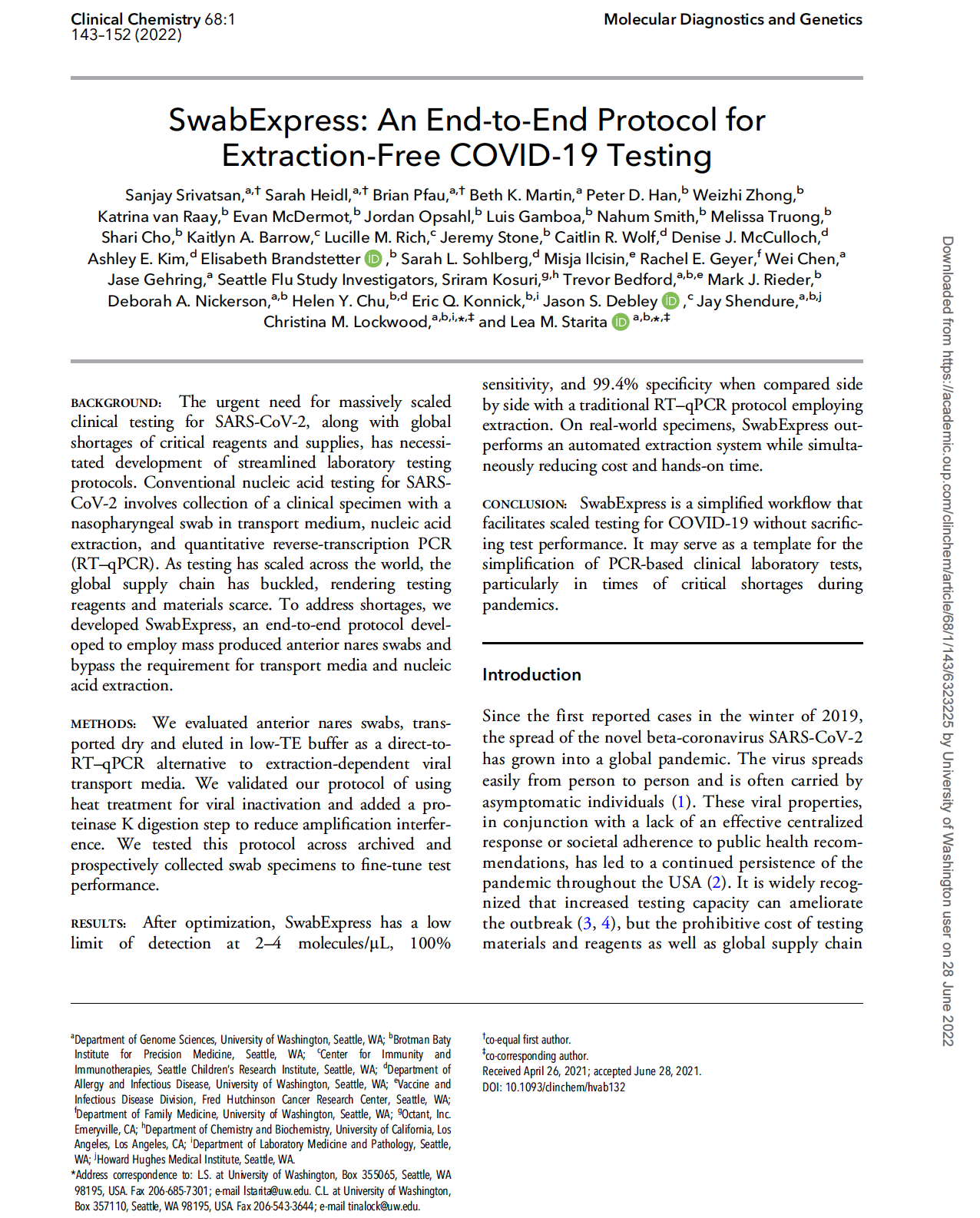
(opens in new tab) Srivatsan et al. SwabExpress: An end-to-end protocol for extraction-free COVID-19 testing.
Clinical Chemistry (2021)
PMID: 32511368 (opens in new tab) | PDF (opens in new tab)
(with Seattle Flu Study)
2020
 (opens in new tab)
(opens in new tab) Bedford, Greninger, Roychoudhury, Starita, Famulare et al. Cryptic transmission of SARS-CoV-2 in Washington state.
Science (2020)
 (opens in new tab)
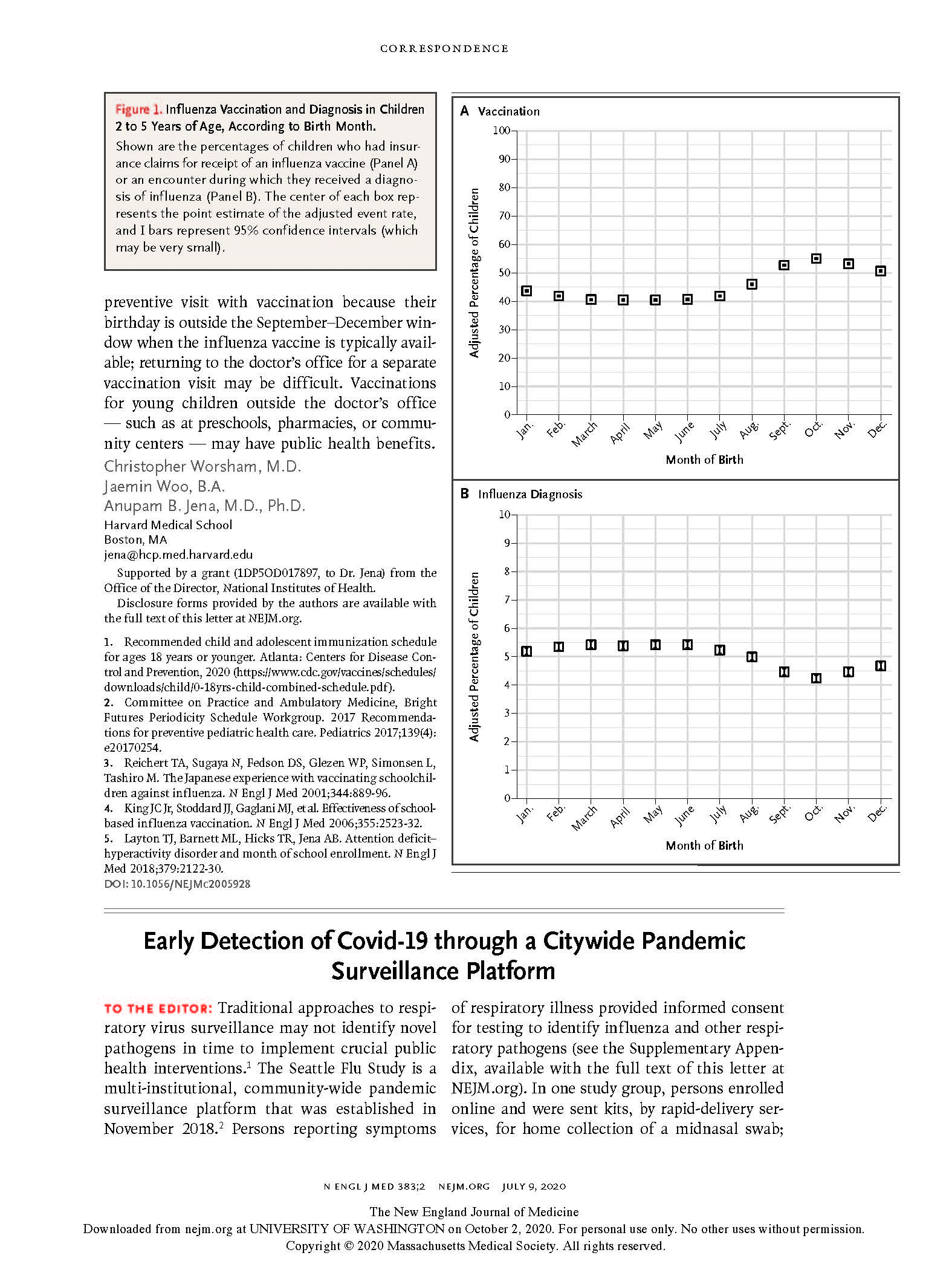
(opens in new tab) Chu et al. Early Detection of Covid-19 through a Citywide Pandemic Surveillance Platform.
NEJM (2020)
 (opens in new tab)
(opens in new tab) Srivatsan, McFaline-Figueroa, Ramani et al. Massively multiplex chemical transcriptomics at single-cell resolution.
Science (2020)
PMID: 31806696 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
2019
 (opens in new tab)
(opens in new tab) Rentzsch et al. CADD: predicting the deleteriousness of variants throughout the human genome.
Nucleic Acids Research (2019)
PMID: 30371827 (opens in new tab) | PDF (opens in new tab)
(with Kircher Lab)
2018
 (opens in new tab)
(opens in new tab) Findlay et al. Accurate classification of BRCA1 variants with saturation genome editing.
Nature (2018)
PMID: 30209399 (opens in new tab) | PDF (opens in new tab)
(with Starita Lab)
 (opens in new tab)
(opens in new tab) Starita et al. A Multiplex Homology-Directed DNA Repair Assay Reveals the Impact of More Than 1,000 BRCA1 Missense Substitution Variants on Protein Function.
AJHG (2018)
PMID: 30219179 (opens in new tab) | PDF (opens in new tab)
(with Parvin Lab)
2017
 (opens in new tab)
(opens in new tab) Starita et al. Variant Interpretation: Functional Assays to the Rescue.
AJHG (2017)
 (opens in new tab)
(opens in new tab) Neveling, Mensenkamp et al. BRCA Testing by Single-Molecule Molecular Inversion Probes.
Clinical Chemistry (2017)
PMID: 27974384 (opens in new tab) | PDF (opens in new tab)
(with Nelen, Hoischen Labs)
2016
 (opens in new tab)
(opens in new tab) Underhill et al. Fragment Length of Circulating Tumor DNA.
PLoS Genetics (2016)
PMID: 27428049 (opens in new tab) | PDF (opens in new tab)
(with Underhill Lab)
 (opens in new tab)
(opens in new tab) Snyder, Kircher et al. Cell-free DNA Comprises an In Vivo Nucleosome Footprint that Informs Its Tissues-Of-Origin.
Cell (2016)
2015
 (opens in new tab)
(opens in new tab) Roach et al. A Year of Infection in the Intensive Care Unit: Prospective Whole Genome Sequencing of Bacterial Clinical Isolates Reveals Cryptic Transmissions and Novel Microbiota.
PLoS Genetics (2015)
PMID: 26230489 (opens in new tab) | PDF (opens in new tab)
(with Salipante Lab)
 (opens in new tab)
(opens in new tab) Starita et al. Massively Parallel Functional Analysis of BRCA1 RING Domain Variants.
Genetics (2015)
PMID: 25823446 (opens in new tab) | PDF (opens in new tab)
(with Fields Lab)
 (opens in new tab)
(opens in new tab) Snyder, Simmons et al. Copy-Number Variation and False Positive Prenatal Aneuploidy Screening Results.
New England Journal of Medicine (2015)
PMID: 25830323 (opens in new tab) | PDF (opens in new tab)
(with Gammill Lab)
 (opens in new tab)
(opens in new tab) Kumar, Ryan et al. Whole genome prediction for preimplantation genetic diagnosis.
Genome Medicine (2015)
2014
 (opens in new tab)
(opens in new tab) Salipante, Roach et al. Large-scale genomic sequencing of extraintestinal pathogenic Escherichia coli strains.
Genome Research (2014)
 (opens in new tab)
(opens in new tab) Kumar et al. Deep sequencing of multiple regions of glial tumors reveals spatial heterogeneity for mutations in clinically relevant genes.
Genome Biology (2014)
PMID: 25608559 (opens in new tab) | PDF (opens in new tab)
(with Rostomily Lab)
 (opens in new tab)
(opens in new tab) Kumar et al. Genome sequencing of idiopathic pulmonary fibrosis in conjunction with a medical school human anatomy course.
PLoS One (2014)
 (opens in new tab)
(opens in new tab) Kircher, Witten et al. A general framework for estimating the relative pathogenicity of human genetic variants.
Nature Genetics (2014)
PMID: 24487276 (opens in new tab) | PDF (opens in new tab)
(with Cooper Lab)
2013
 (opens in new tab)
(opens in new tab) Snyder et al. Noninvasive fetal genome sequencing: a primer.
Prenatal Diagnosis (2013)
2012
 (opens in new tab)
(opens in new tab) Kitzman et al. Noninvasive whole-genome sequencing of a human fetus.
Science Translational Medicine (2012)
2011
 (opens in new tab)
(opens in new tab) Cooper, Shendure. Needles in stacks of needles: finding disease-causal variants in a wealth of genomic data.
Nature Reviews Genetics (2011)
2009
 (opens in new tab)
(opens in new tab) Vasta V et al. Next generation sequence analysis for mitochondrial disorders.
Genome Medicine (2009)
PMID: 19852779 (opens in new tab) | PDF (opens in new tab)
(with Hahn Lab)

