Developing New Molecular Methods
(selected publications)
2026
 (opens in new tab)
(opens in new tab) Lalanne, Chau et al. Pool-packaged AAV libraries exhibit extensive length-dependent and homology-dependent chimerism.
Nature Biotechnology (2026)
 (opens in new tab)
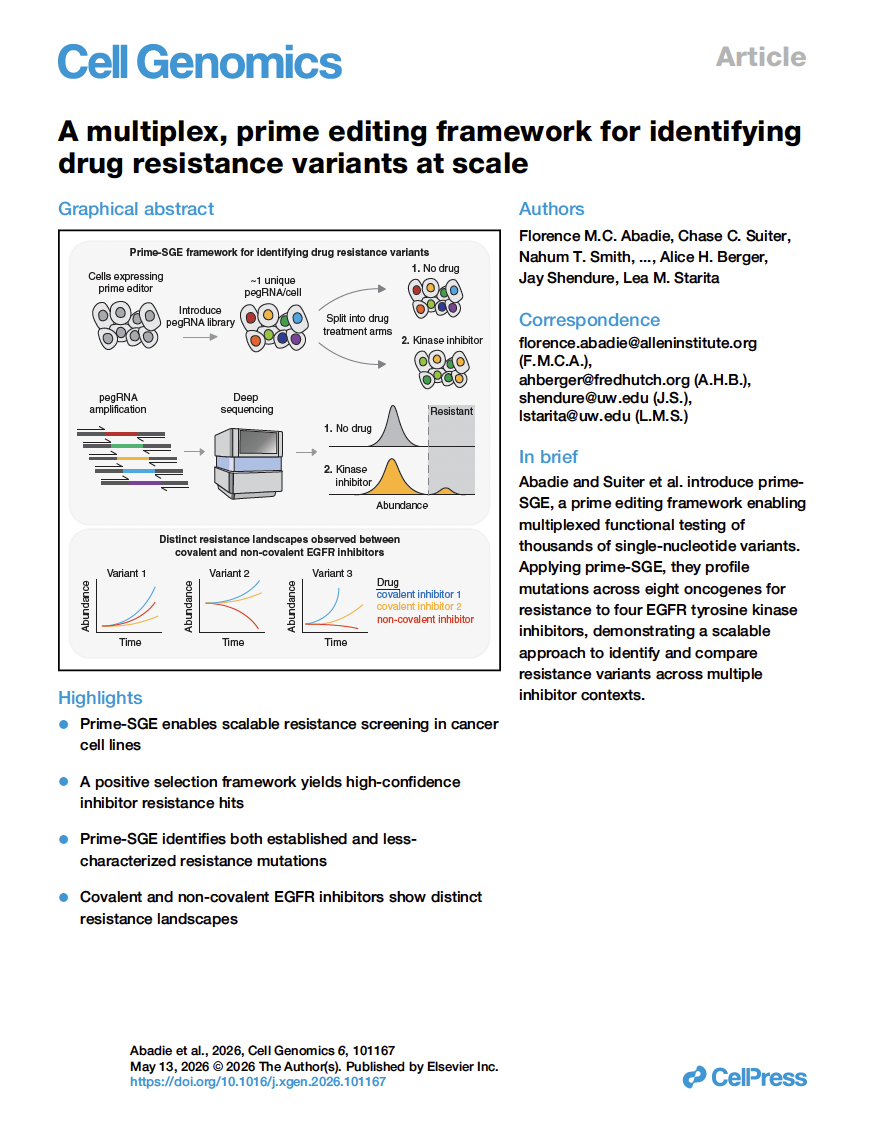
(opens in new tab) Abadie, Suiter et al. A multiplex, prime editing framework for identifying drug resistance variants at scale.
Cell Genomics (2026)
 (opens in new tab)
(opens in new tab) Nathans, McDiarmid et al. Multichannel genomic recording of biological information with ENGRAM.
Nature Protocols (2026)
2025
 (opens in new tab)
(opens in new tab) McDiarmid, Taylor et al. A parts list of promoters and gRNA scaffolds for mammalian genome engineering and molecular recording.
Nature Biotechnology (2025)
 (opens in new tab)
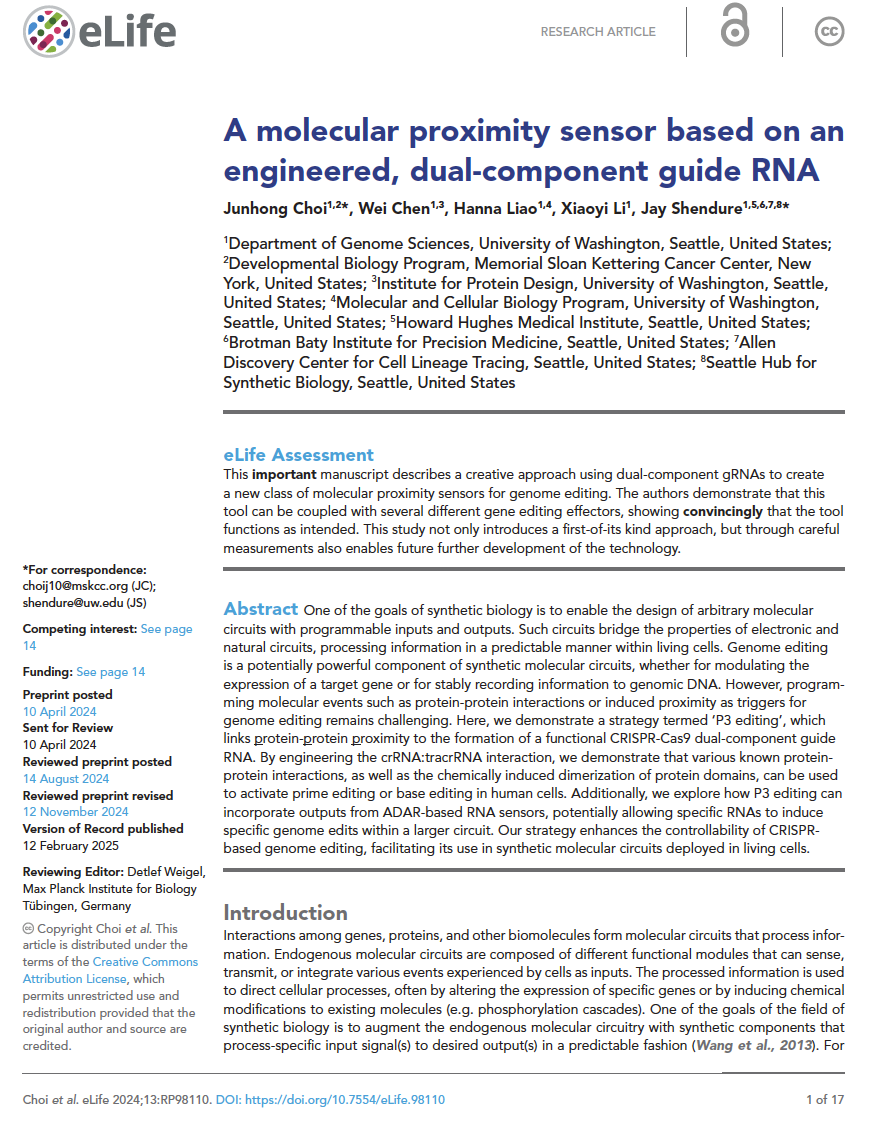
(opens in new tab) Choi et al. A molecular proximity sensor based on an engineered, dual-component guide RNA.
eLife (2025)
2024
 (opens in new tab)
(opens in new tab) Chen, Choi et al. Symbolic recording of signalling and cis-regulatory element activity to DNA.
Nature (2024)
 (opens in new tab)
(opens in new tab) Liao, Choi, Shendure. Molecular recording using DNA Typewriter.
Nature Protocols (2024)
 (opens in new tab)
(opens in new tab) Li et al. Chromatin context-dependent regulation and epigenetic manipulation of prime editing.
Cell (2024)
2022
 (opens in new tab)
(opens in new tab) Martin et al. Optimized single-nucleus transcriptional profiling by combinatorial indexing.
Nature Protocols (2022)
 (opens in new tab)
(opens in new tab) Choi et al. A time-resolved, multi-symbol molecular recorder via sequential genome editing.
Nature (2022)
 (opens in new tab)
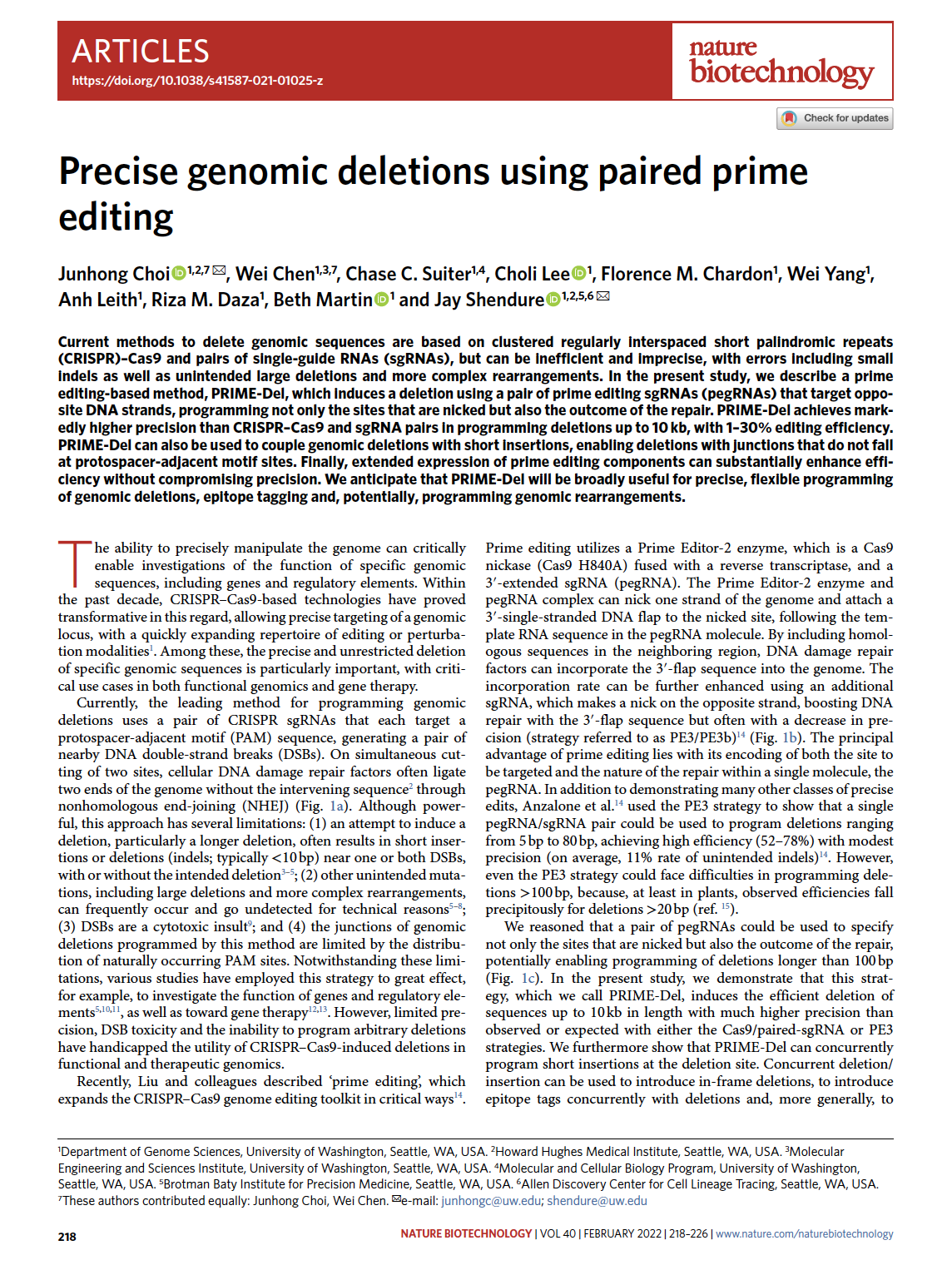
(opens in new tab) Choi, Chen et al. Precise genomic deletions using paired prime editing.
Nature Biotechnology (2022)
2021
.png) (opens in new tab)
(opens in new tab) Srivatsan, Regier et al. Embryo-scale, single-cell spatial transcriptomics.
Science (2021)
PMID: 34210887 (opens in new tab) | PDF (opens in new tab)
(with Trapnell and Stevens Labs)
.png) (opens in new tab)
(opens in new tab) Simeonov et al. Single-cell lineage tracing of metastatic cancer reveals selection of hybrid EMT states.
Cancer Cell (2021)
PMID: 34115987 (opens in new tab) | PDF (opens in new tab)
(with Lengner and McKenna Labs)
2020
 (opens in new tab)
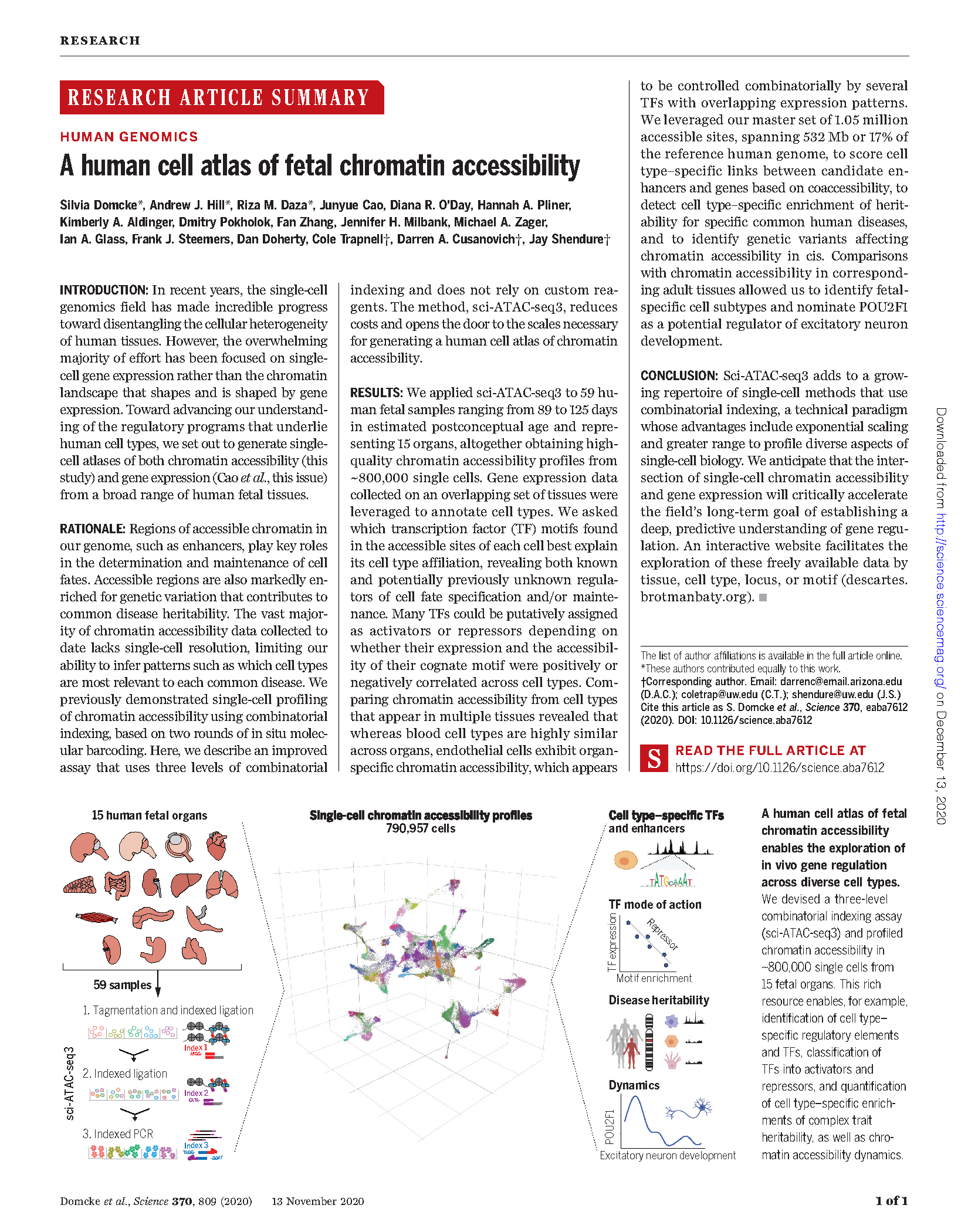
(opens in new tab) Domcke, Hill, Cusanovich, Daza et al. A human cell atlas of fetal chromatin accessibility.
Science (2020)
PMID: 33184180 (opens in new tab) | PDF (opens in new tab)
(with Cusanovich and Trapnell Labs)
 (opens in new tab)
(opens in new tab) Cao et al. Sci-fate characterizes the dynamics of gene expression in single cells.
Nature Biotechnology (2020)
 (opens in new tab)
(opens in new tab) Srivatsan et al. Massively multiplex chemical transcriptomics at single-cell resolution.
Science (2020)
PMID: 31806696 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
2019
 (opens in new tab)
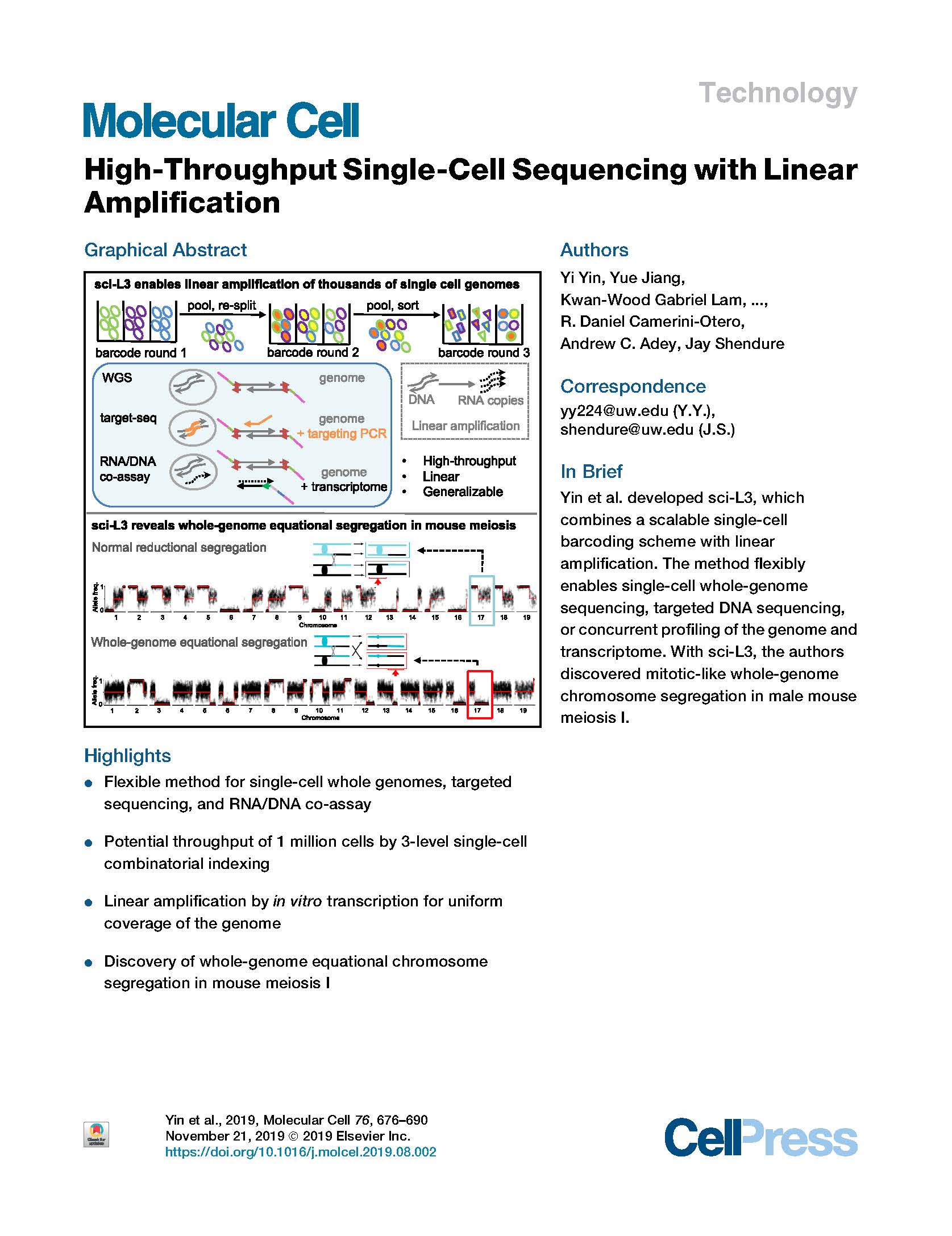
(opens in new tab) Yin et al. High-Throughput Single-Cell Sequencing with Linear Amplification.
Molecular Cell (2019)
 (opens in new tab)
(opens in new tab) Alexander et al. Concurrent genome and epigenome editing by CRISPR-mediated sequence replacement.
BMC Biology (2019)
 (opens in new tab)
(opens in new tab) Pliner et al. Supervised classification enables rapid annotation of cell atlases.
Nature Methods (2019)
PMID: 31501545 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
(opens in new tab) Kircher, Xiong, Martin, Schubach et al. Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution.
Nature Communications (2019)
PMID: 31395865 (opens in new tab) | PDF (opens in new tab)
(with Ahituv and Kircher Labs)
 (opens in new tab)
(opens in new tab) Chen et al. Massively parallel profiling and predictive modeling of the outcomes of CRISPR/Cas9-mediated double-strand break repair.
Nucleic Acids Research (2019)
 (opens in new tab)
(opens in new tab) Kim et al. A combination of transcription factors mediates inducible interchromosomal contacts.
eLife (2019)
 (opens in new tab)
(opens in new tab) Ramani et al. High Sensitivity Profiling of Chromatin Structure by MNase-SSP.
Cell Reports (2019)
 (opens in new tab)
(opens in new tab) Cao, Spielmann et al. The single-cell transcriptional landscape of mammalian organogenesis.
Nature (2019)
 (opens in new tab)
(opens in new tab) Gasperini et al. A Genome-wide Framework for Mapping Gene Regulation via Cellular Genetic Screens.
Cell (2019)
2018
 (opens in new tab)
(opens in new tab) Cao et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells.
Science (2018)
PMID: 30166440 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
(opens in new tab) Matreyek, Starita et al. Multiplex assessment of protein variant abundance by massively parallel sequencing.
Nature Genetics (2018)
PMID: 29785012 (opens in new tab) | PDF (opens in new tab)
(with Fowler Lab)
 (opens in new tab)
(opens in new tab) Hill, McFaline-Figueroa et al. On the design of CRISPR-based single-cell molecular screens.
Nature Methods (2018)
PMID: 29457792 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
2017
 (opens in new tab)
(opens in new tab) Cao, Packer et al. Comprehensive single cell transcriptional profiling of a multicellular organism by combinatorial indexing.
Science (2017)
PMID: 28818938 (opens in new tab) | PDF (opens in new tab)
(with Trapnell and Waterston Labs)
 (opens in new tab)
(opens in new tab) Gasperini, Findlay et al. CRISPR/Cas9-Mediated Scanning for Regulatory Elements Required for HPRT1 Expression via Thousands of Large, Programmed Genomic Deletions.
AJHG (2017)
2016
 (opens in new tab)
(opens in new tab) Ramani, Cusanovich et al. Mapping 3D genome architecture through in situ DNase Hi-C.
Nature Protocols (2016)
 (opens in new tab)
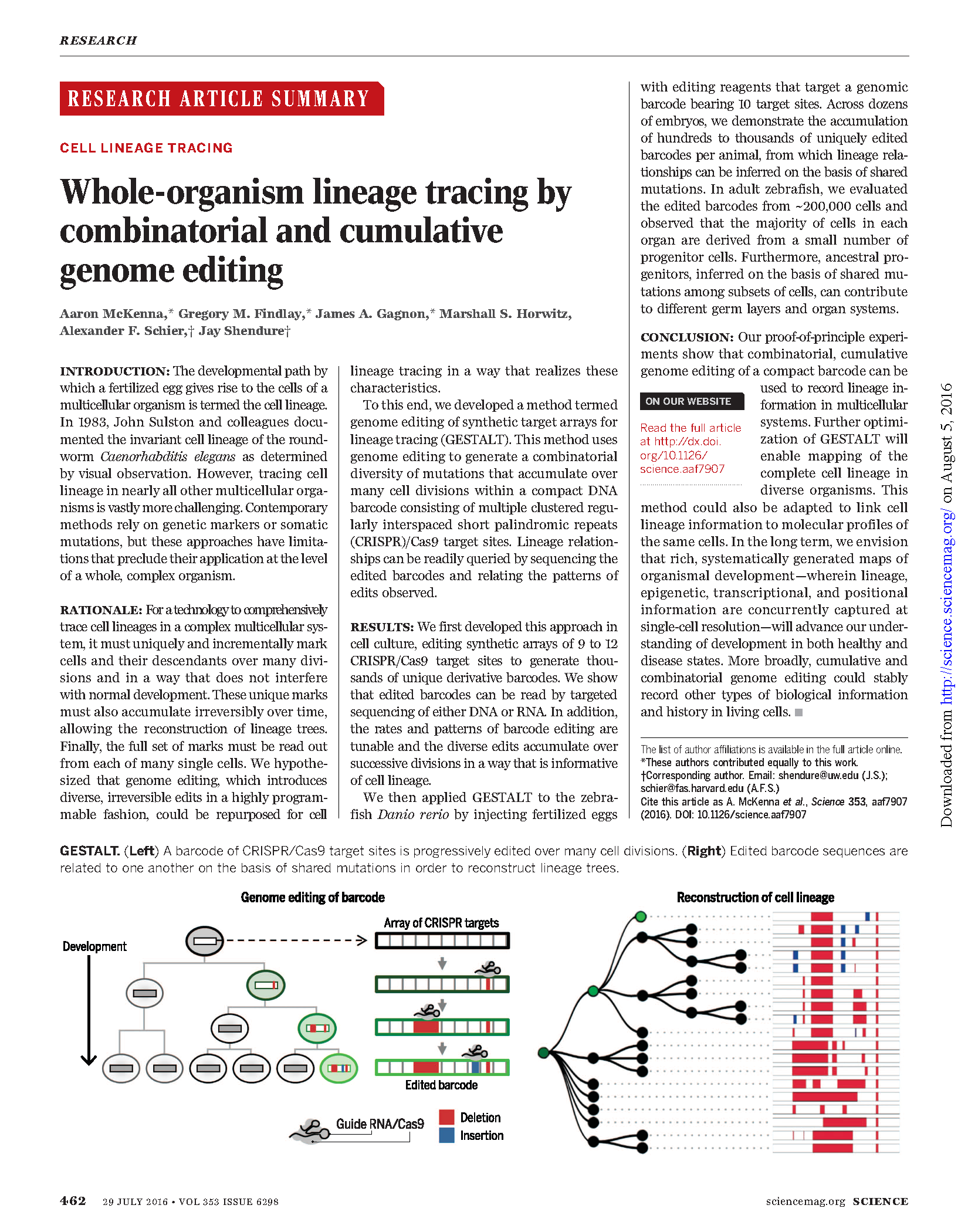
(opens in new tab) McKenna, Findlay, Gagnon et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing.
Science (2016)
PMID: 27229144 (opens in new tab) | PDF (opens in new tab)
(with Schier Lab)
2015
 (opens in new tab)
(opens in new tab) Klein et al. Multiplex pairwise assembly of array-derived DNA oligonucleotides.
Nucleic Acids Research (2015)
 (opens in new tab)
(opens in new tab) Ramani et al. High-throughput determination of RNA structure by proximity ligation.
Nature Biotechnology (2015)
 (opens in new tab)
(opens in new tab) Cusanovich et al. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing.
Science (2015)
PMID: 25953818 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
(opens in new tab) Kitzman, Starita et al. Massively parallel single-amino-acid mutagenesis.
Nature Methods (2015)
PMID: 25559584 (opens in new tab) | PDF (opens in new tab)
(with Fields Lab)
 (opens in new tab)
(opens in new tab) Snyder et al. Haplotype-resolved genome sequencing: experimental methods and applications.
Nature Reviews Genetics (2015)
2014
 (opens in new tab)
(opens in new tab) Boyle et al. MIPgen: optimized modeling and design of molecular inversion probes for targeted resequencing.
Bioinformatics (2014)
2013
 (opens in new tab)
(opens in new tab) Hiatt et al. Single molecule molecular inversion probes for targeted, high-accuracy detection of low-frequency variation.
Genome Research (2013)
2012
 (opens in new tab)
(opens in new tab) Shendure, Lieberman-Aiden. The expanding scope of DNA sequencing.
Nature Biotechnology (2012)
 (opens in new tab)
(opens in new tab) Adey, Shendure. Ultra-low-input, tagmentation-based whole-genome bisulfite sequencing.
Genome Research (2012)
 (opens in new tab)
(opens in new tab) Schwartz et al. Accurate gene synthesis with tag-directed retrieval of sequence-verified DNA molecules.
Nature Methods (2012)
2010
 (opens in new tab)
(opens in new tab) Adey, Morrison, Asan, Xun et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high-density in vitro transposition.
Genome Biology (2010)
PMID: 21143862 (opens in new tab) | PDF (opens in new tab)
(with Zhang Lab)
 (opens in new tab)
(opens in new tab) Hiatt, Patwardhan et al. Parallel, tag-directed assembly of locally derived short sequence reads.
Nature Methods (2010)
2009
 (opens in new tab)
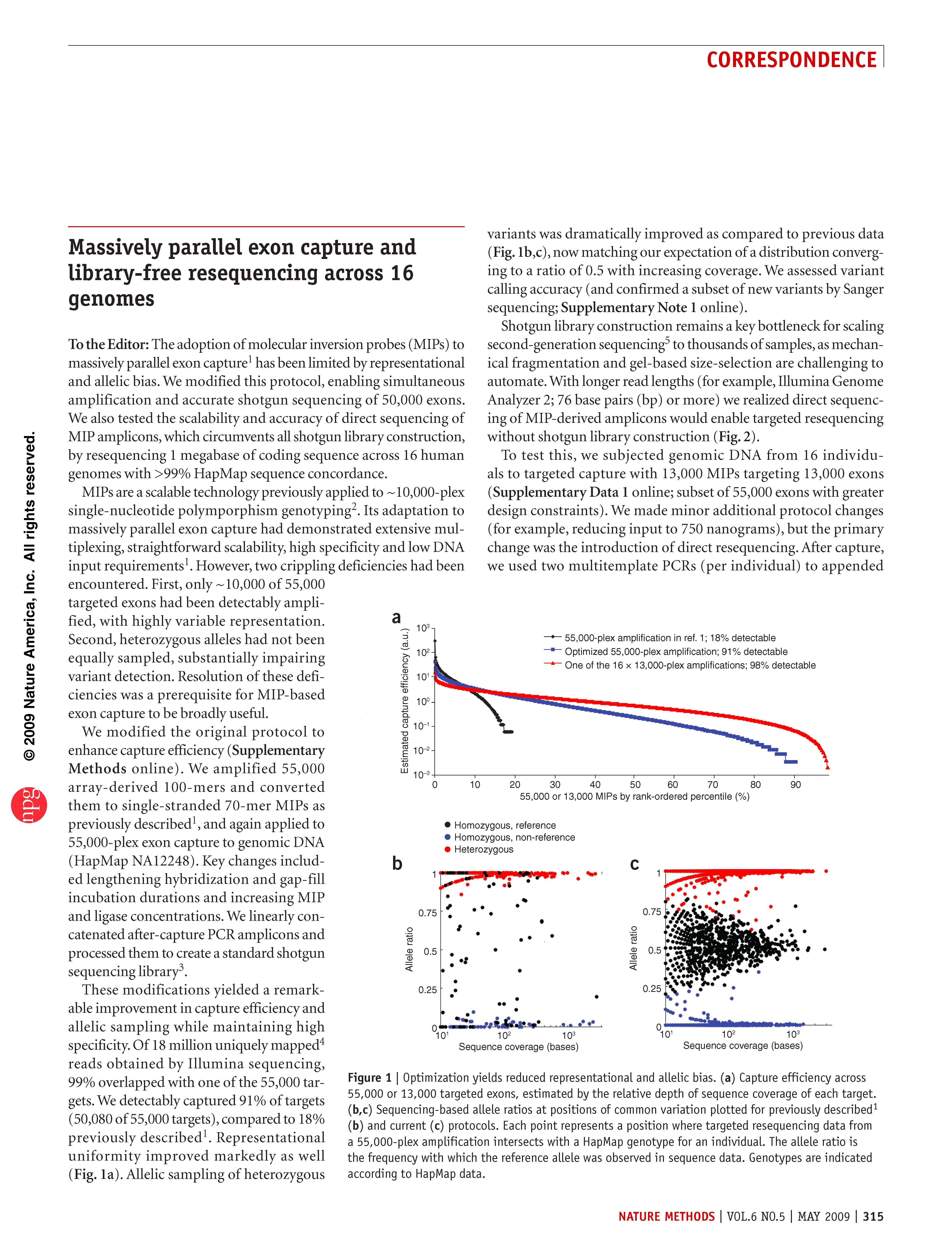
(opens in new tab) Turner et al. Massively parallel exon capture and library-free resequencing across 16 genomes.
Nature Methods (2009)
2007
 (opens in new tab)
(opens in new tab) Porreca, Zhang et al. Multiplex Amplification of Large Sets of Human Exons.
Nature Methods (2007)
PMID: 17934468 (opens in new tab) | PDF (opens in new tab)
(in Church Lab)
