Featured Research Articles
(work led or co-led by our lab)
2026
 (opens in new tab)
(opens in new tab) Lalanne, Huynh et al. Pool-packaged AAV libraries exhibit extensive length-dependent and homology-dependent chimerism
 (opens in new tab)
(opens in new tab) Garge et al. The proteomic landscape and temporal dynamics of human and mouse gastruloid development
 (opens in new tab)
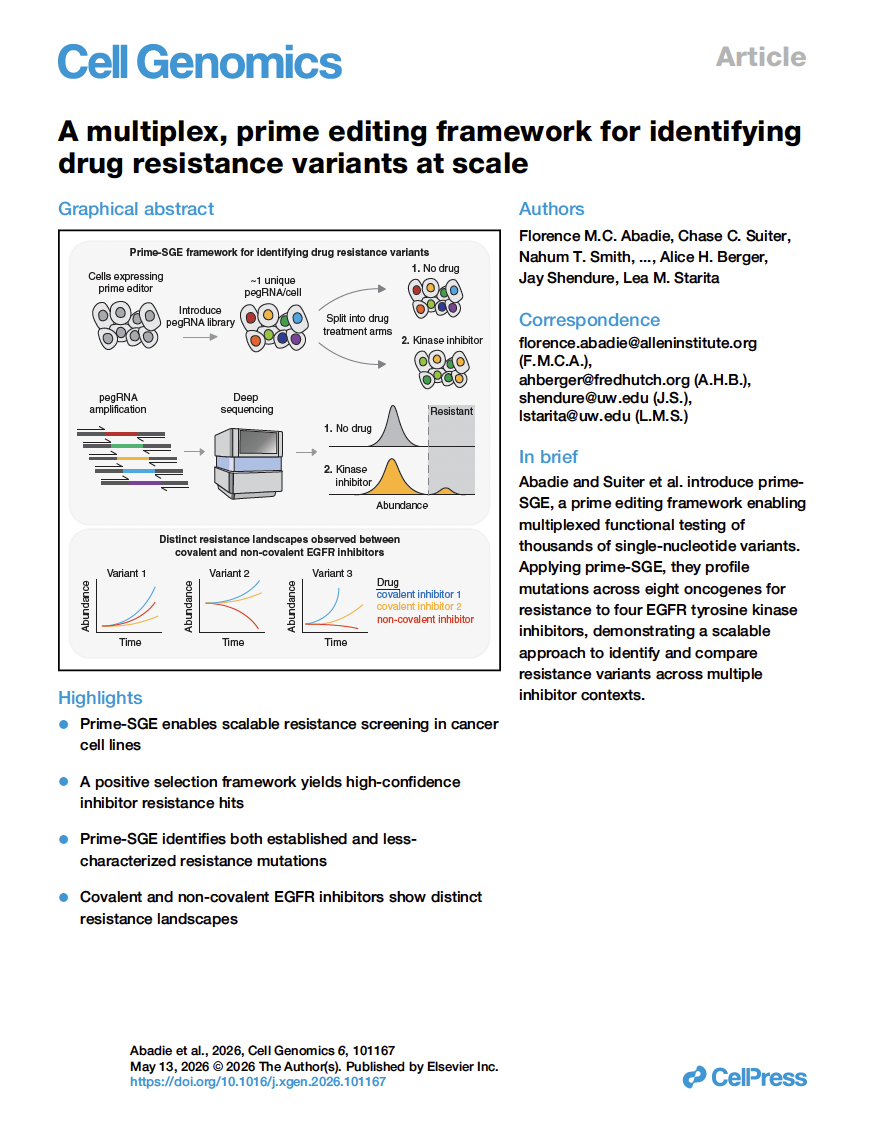
(opens in new tab) Abadie, Suiter et al. A multiplex, prime editing framework for identifying drug resistance variants at scale
2025
 (opens in new tab)
(opens in new tab) McDiarmid, Taylor et al. A parts list of promoters and gRNA scaffolds for mammalian genome engineering and molecular recording
 (opens in new tab)
(opens in new tab) Choi et al. A molecular proximity sensor based on an engineered, dual-component guide RNA
 (opens in new tab)
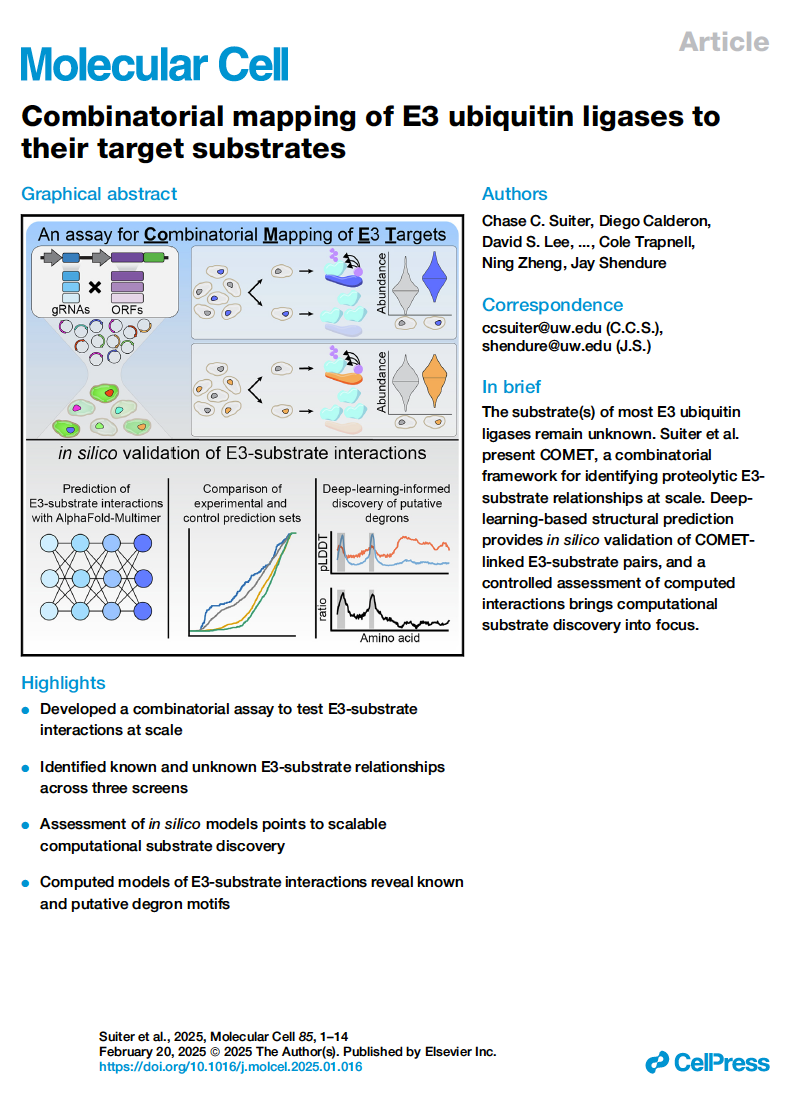
(opens in new tab) Suiter et al. Combinatorial mapping of E3 ubiquitin ligases to their target substrates
 (opens in new tab)
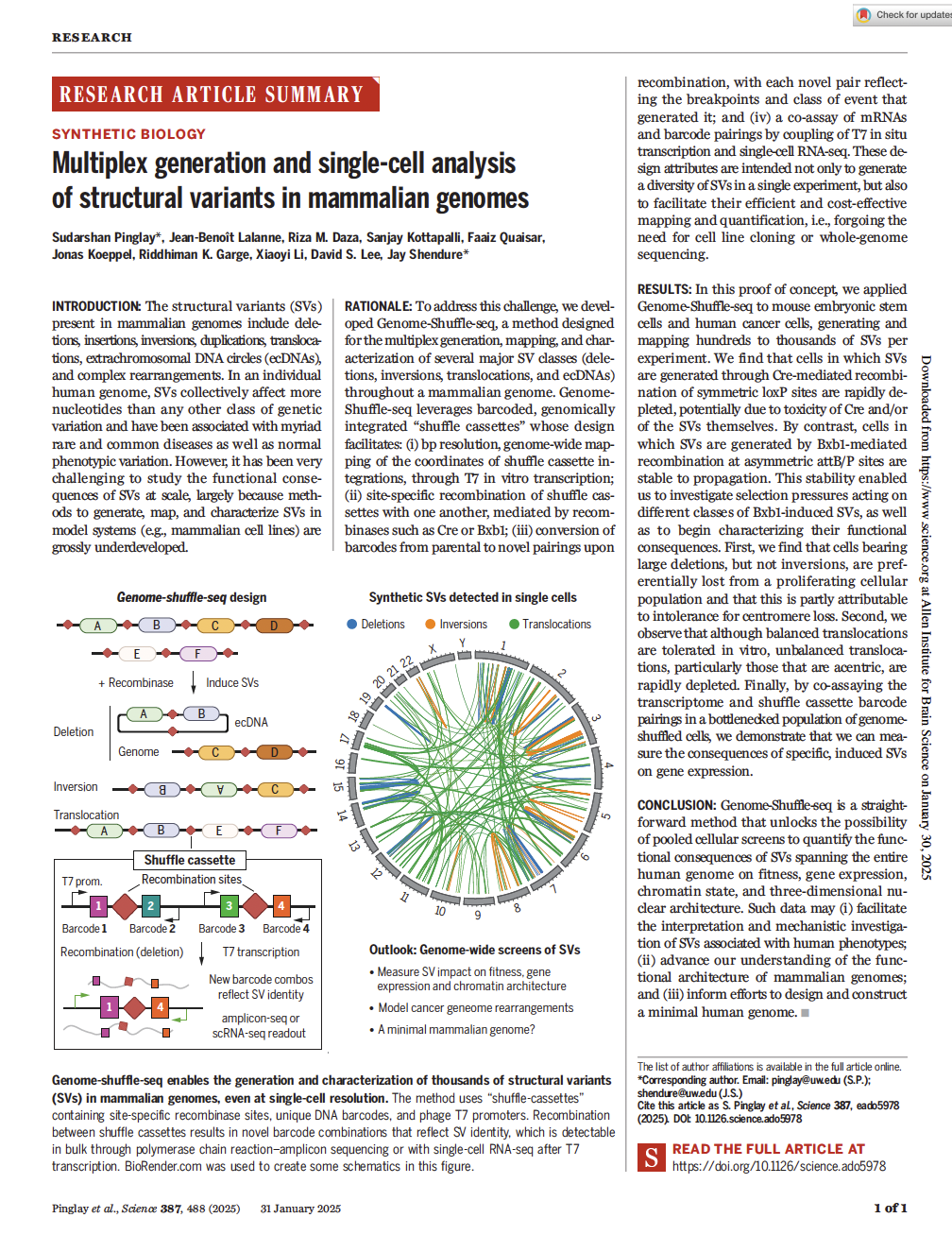
(opens in new tab) Pinglay et al. Multiplex generation and single-cell analysis of structural variants in mammalian genomess
 (opens in new tab)
(opens in new tab) Agarwal, Inoue et al. Massively parallel characterization of transcriptional regulatory elements
2024
 (opens in new tab)
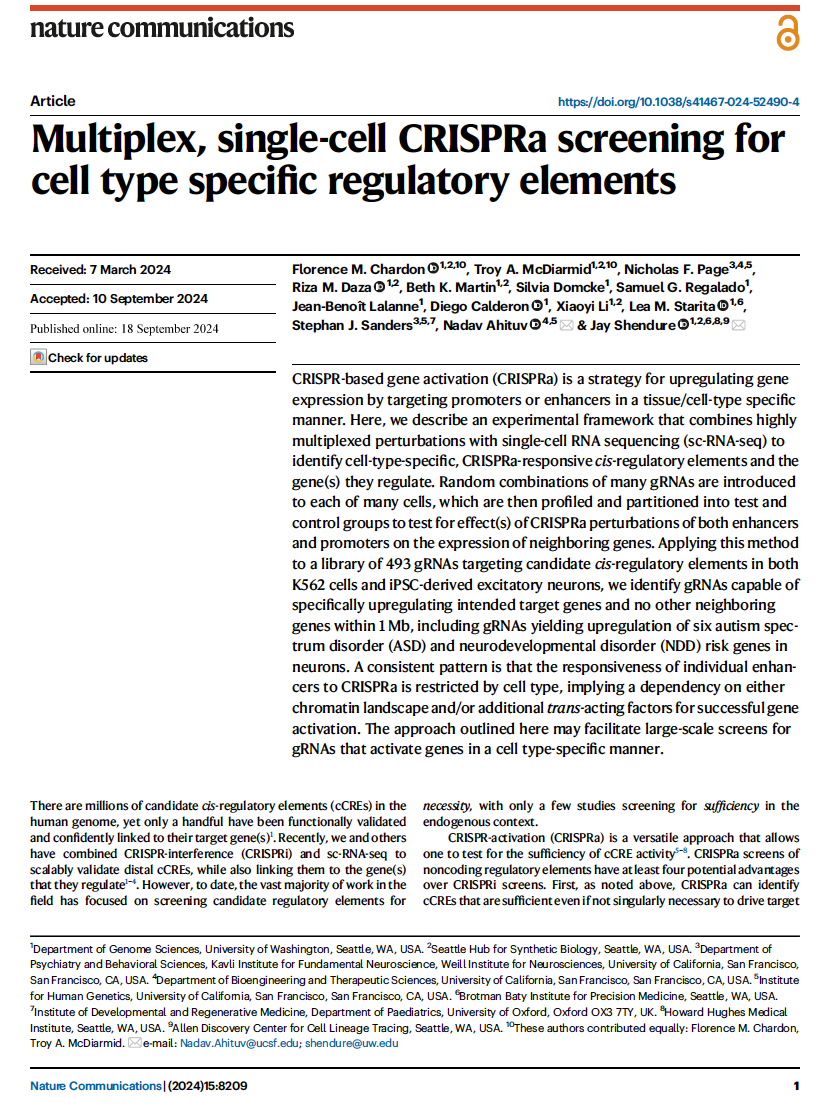
(opens in new tab) Chardon, McDiarmid et al. Multiplex, single-cell CRISPRa screening for cell type specific regulatory elements
 (opens in new tab)
(opens in new tab) Hamazaki, Yang et al. Retinoic acid induces human gastruloids with posterior embryo-like structures
 (opens in new tab)
(opens in new tab) Chen, Choi et al. Symbolic recording of signalling and cis-regulatory element activity to DNA
 (opens in new tab)
(opens in new tab) Lalanne, Regalado et al. Multiplex profiling of developmental cis-regulatory elements with quantitative single-cell expression reporters
 (opens in new tab)
(opens in new tab) Li et al. Chromatin context-dependent regulation and epigenetic manipulation of prime editing
 (opens in new tab)
(opens in new tab) Qiu, Martin, Welsh et al. A single-cell time-lapse of mouse prenatal development from gastrula to birth
 (opens in new tab)
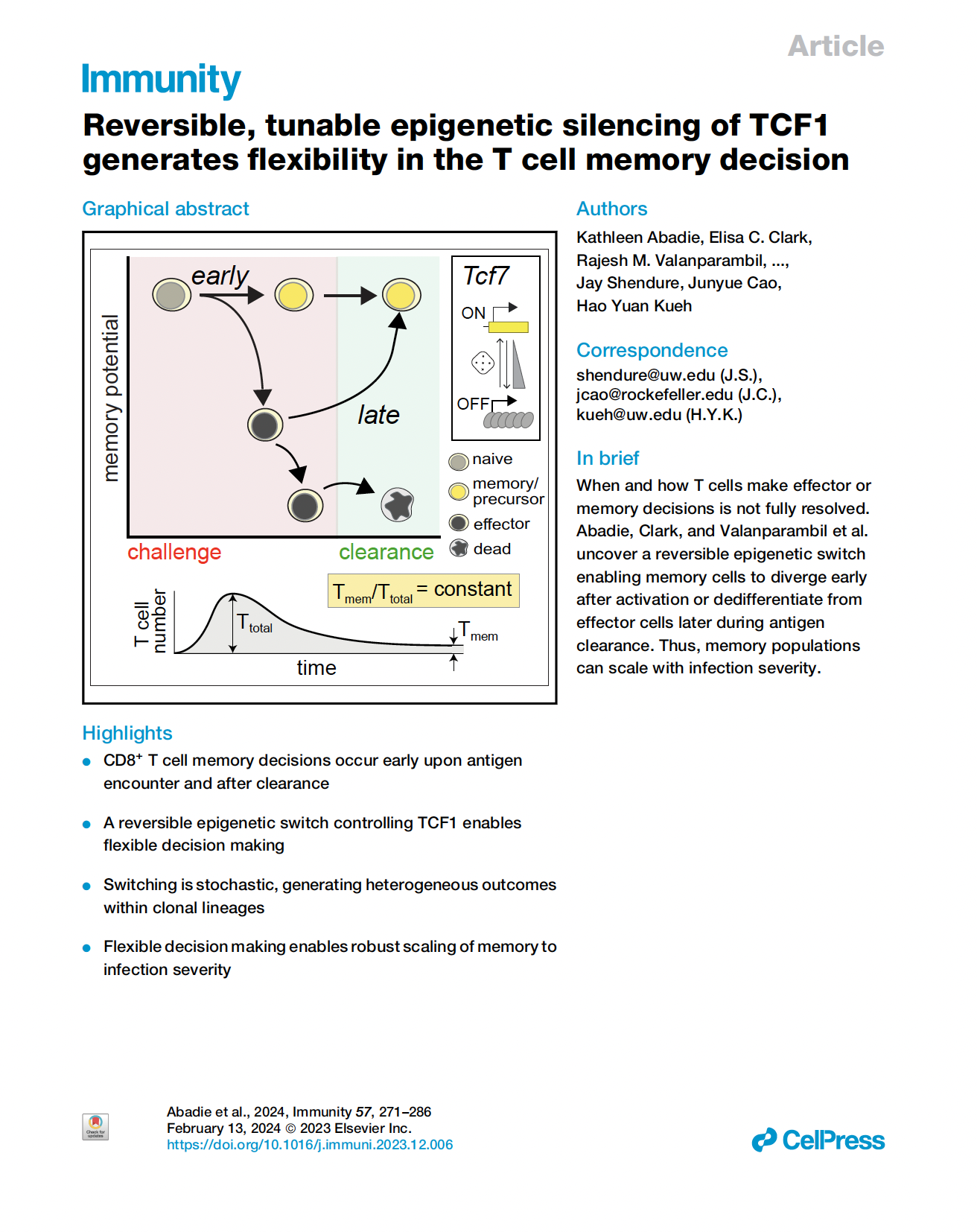
(opens in new tab) Abadie, Clark, Valanparambil et al. Reversible, tunable epigenetic silencing of TCF-1 generates flexibility in the T cell memory decision
2023
 (opens in new tab)
(opens in new tab) Huang, Henck, Qiu et al. Single-cell, whole embryo phenotyping of mammalian developmental disorders
 (opens in new tab)
(opens in new tab) Chiou, Huang et al. A single-cell multi-omic atlas spanning the adult rhesus macaque brain
2022
 (opens in new tab)
(opens in new tab) Martin et al. Optimized single-nucleus transcriptional profiling by combinatorial indexing
 (opens in new tab)
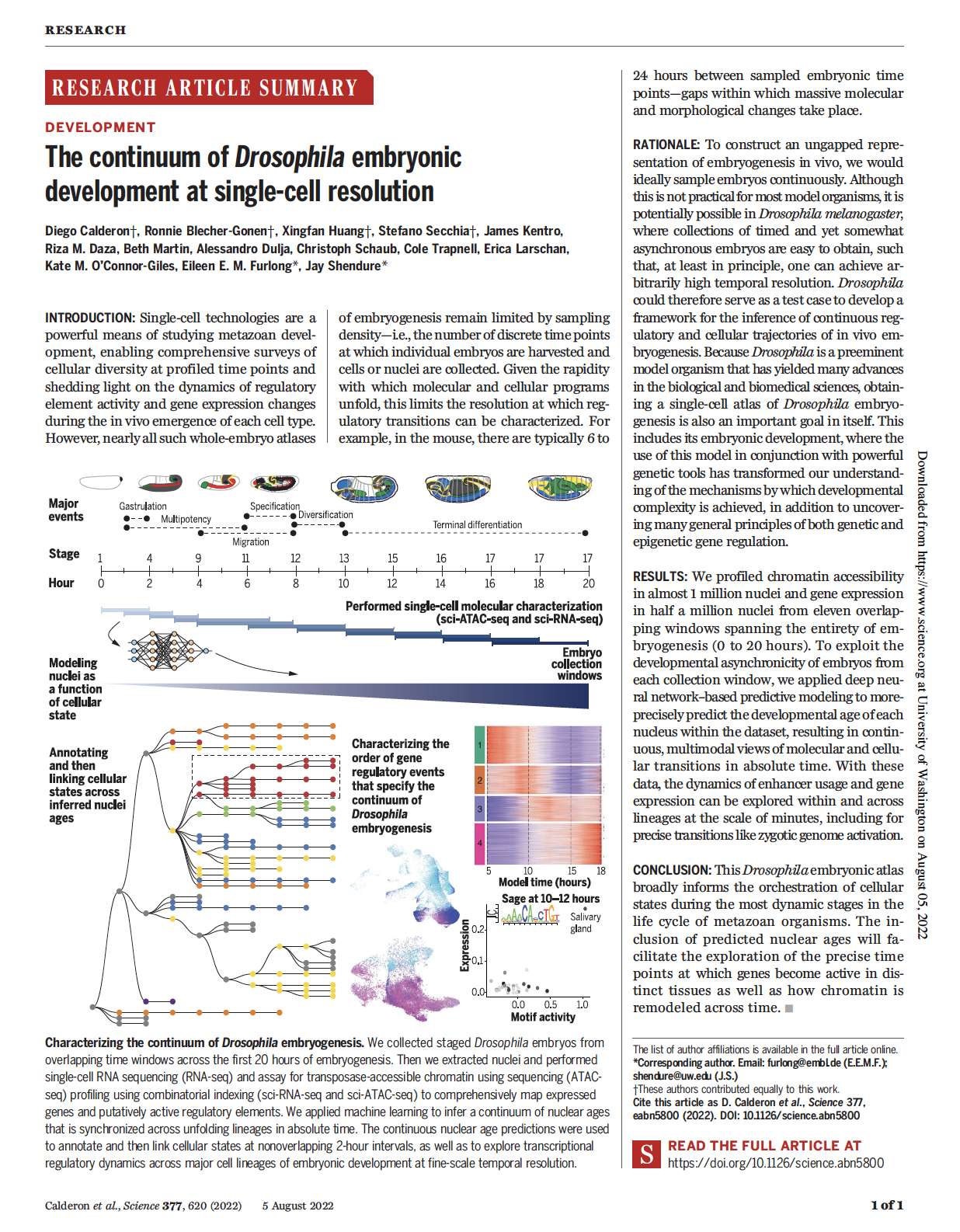
(opens in new tab) Calderon, Huang et al. The continuum of Drosophila embryonic development at single-cell resolution
 (opens in new tab)
(opens in new tab) Choi et al. A time-resolved, multi-symbol molecular recorder via sequential genome editing
 (opens in new tab)
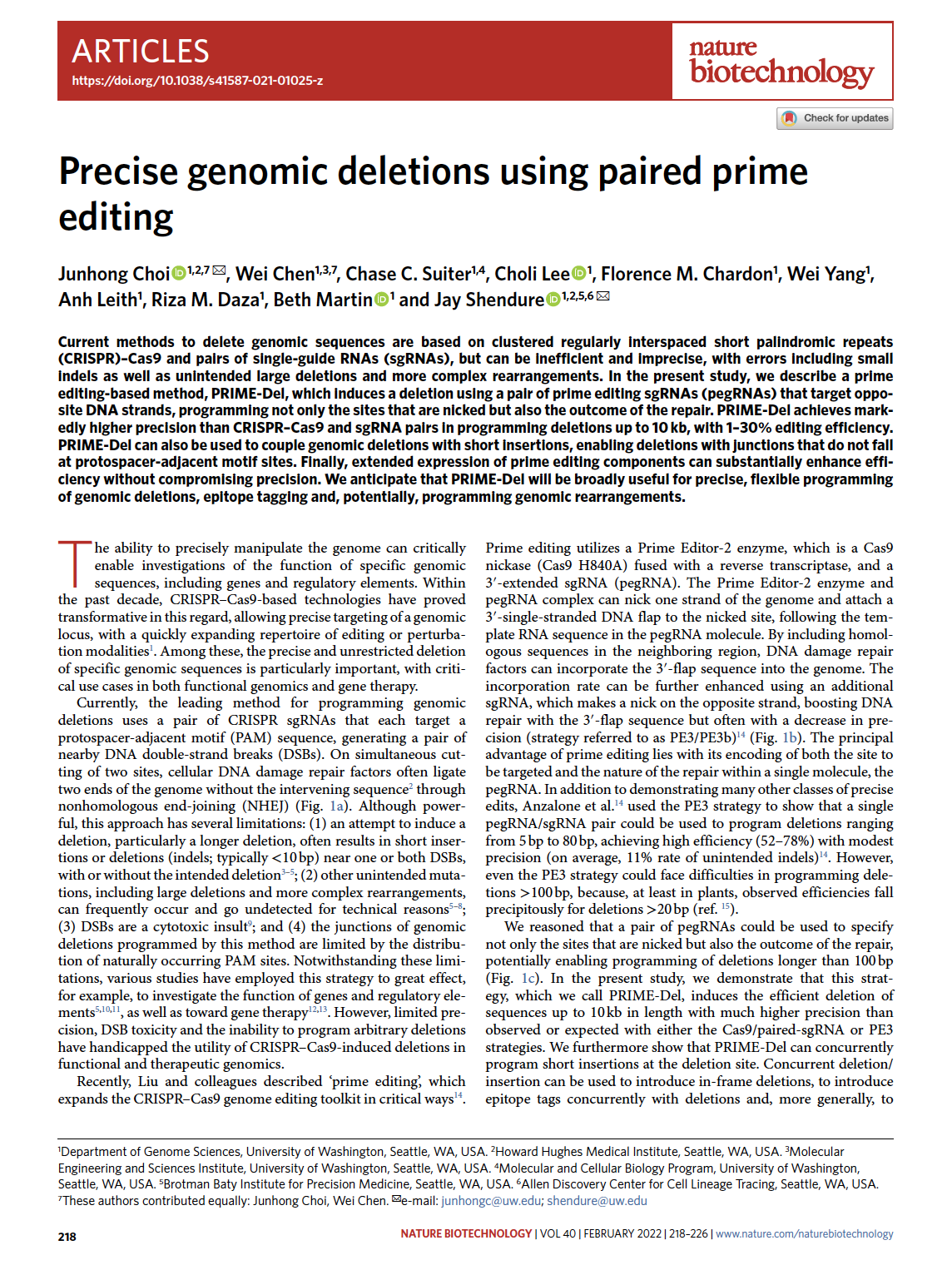
(opens in new tab) Choi, Chen et al. Precise genomic deletions using paired prime editing
 (opens in new tab)
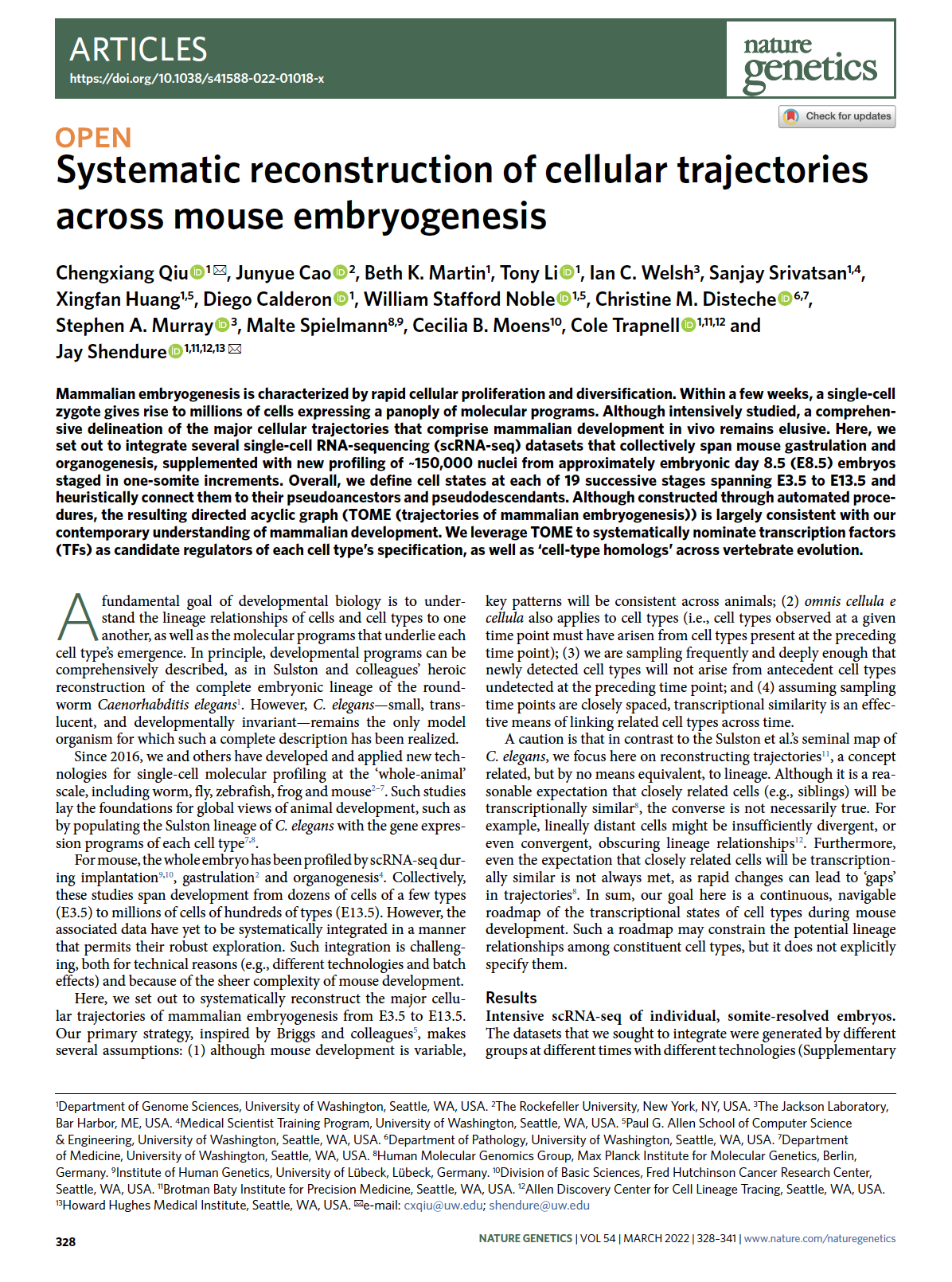
(opens in new tab) Qiu et al. Systematic reconstruction of cellular trajectories across mouse embryogenesis
2021
 (opens in new tab)
(opens in new tab) Agarwal, Darwin-Lopez et al. The landscape of alternative polyadenylation in single cells of the developing mouse embryo
.png) (opens in new tab)
(opens in new tab) Srivatsan, Regier et al. Embryo-scale, single-cell spatial transcriptomics
 (opens in new tab)
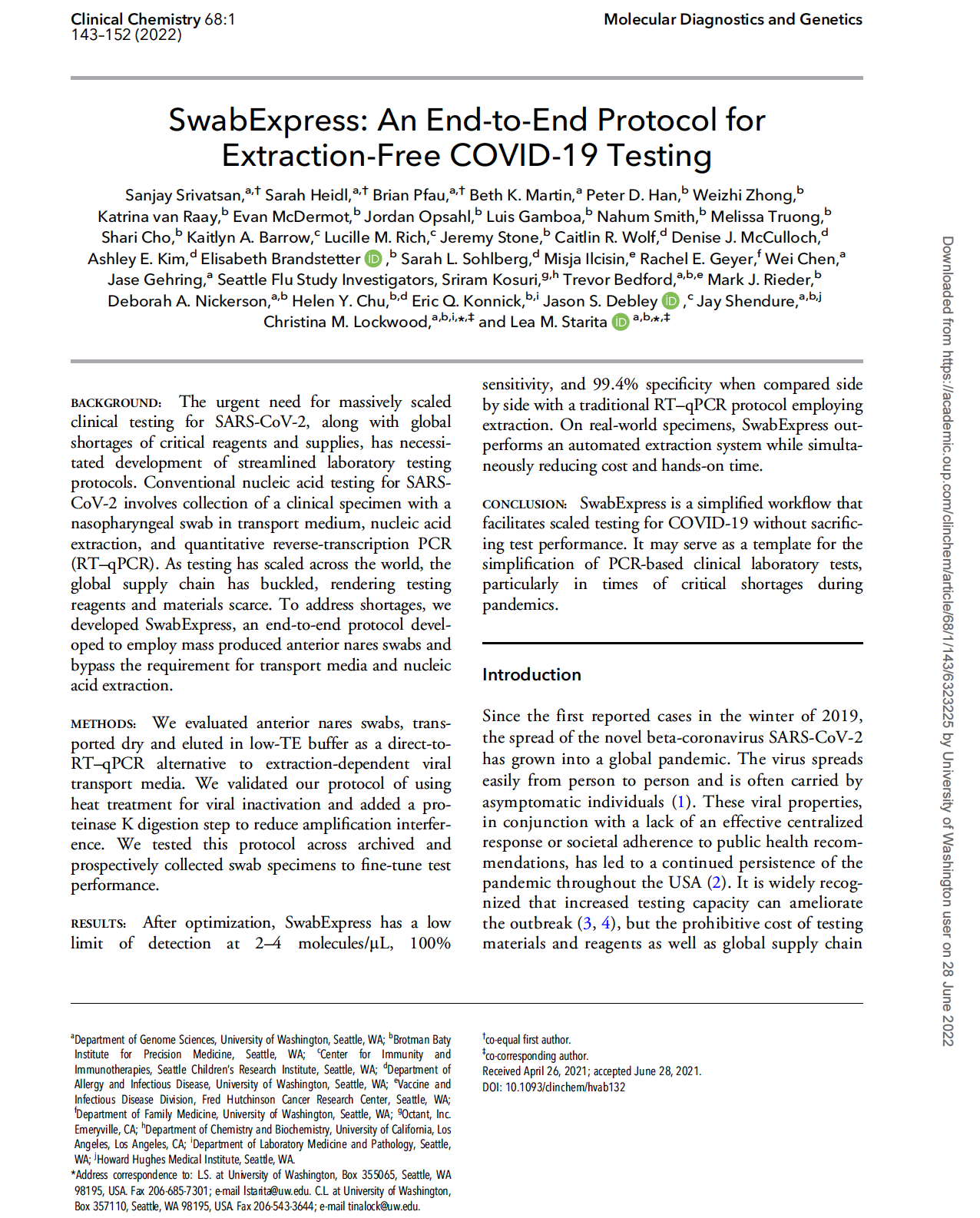
(opens in new tab) Srivatsan et al. SwabExpress: An end-to-end protocol for extraction-free COVID-19 testing
.png) (opens in new tab)
(opens in new tab) Simeonov et al. Single-cell lineage tracing of metastatic cancer reveals selection of hybrid EMT states
2020
 (opens in new tab)
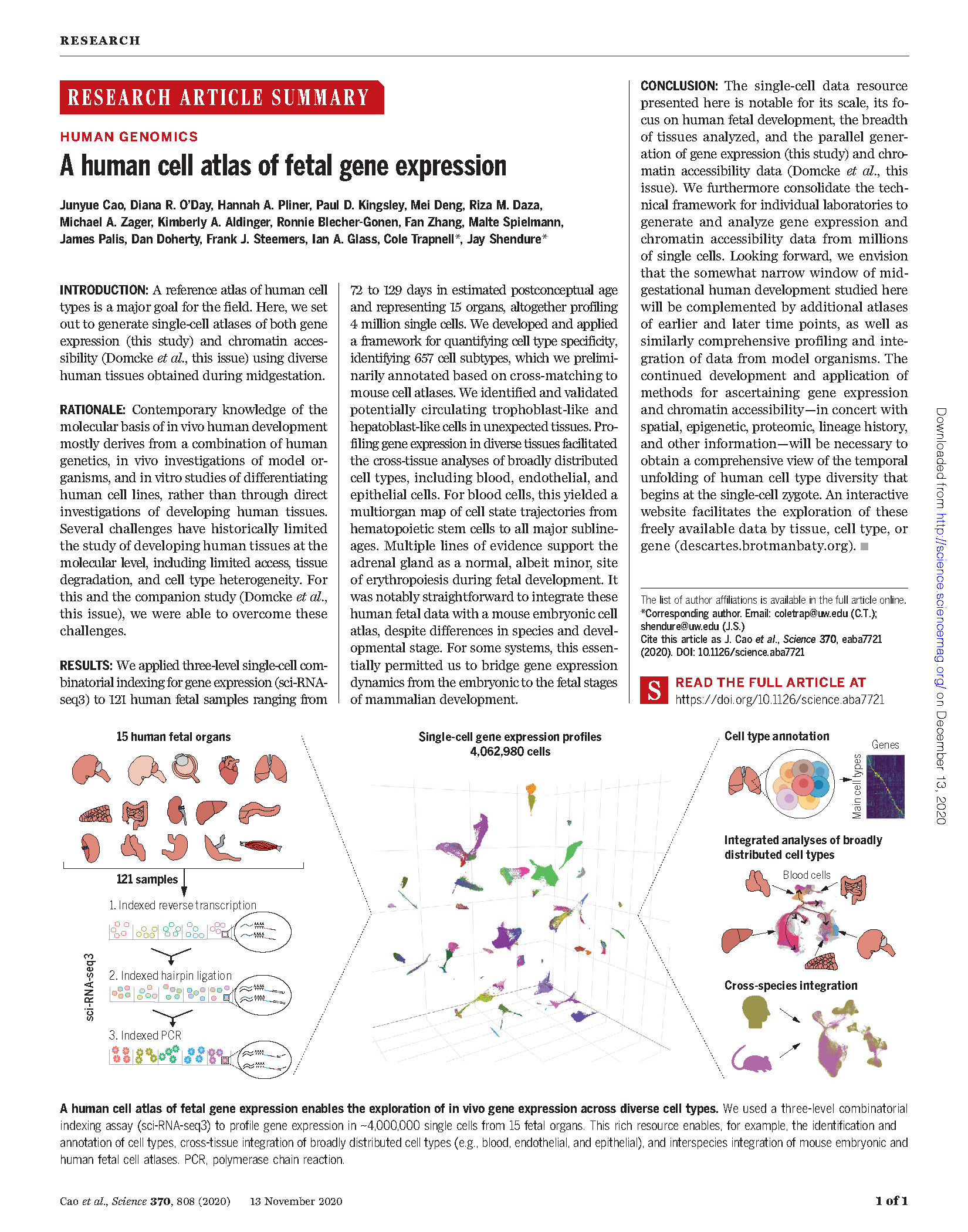
(opens in new tab) Cao et al. A human cell atlas of fetal gene expression
 (opens in new tab)
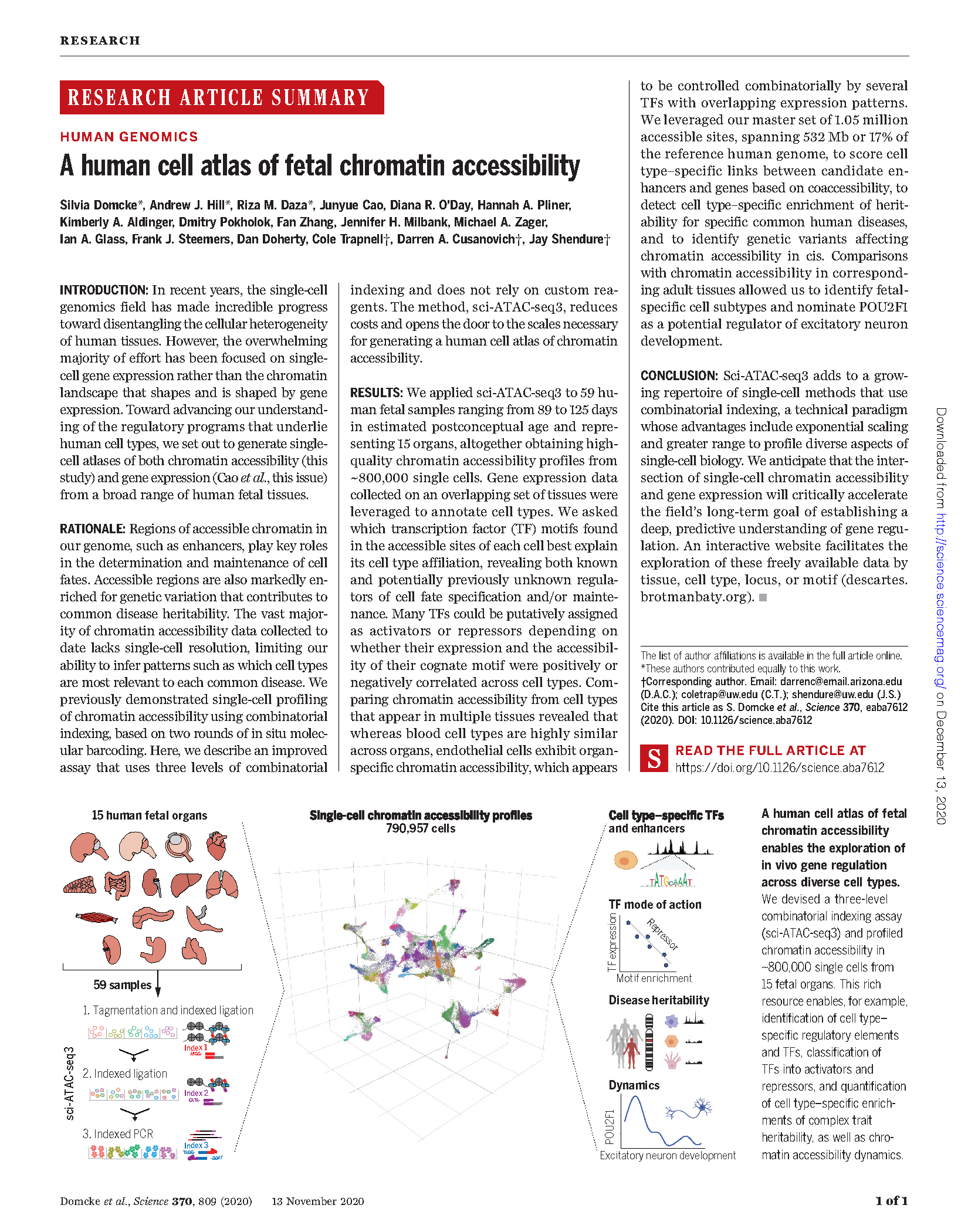
(opens in new tab) Domcke, Hill, Daza et al. A human cell atlas of fetal chromatin accessibility
 (opens in new tab)
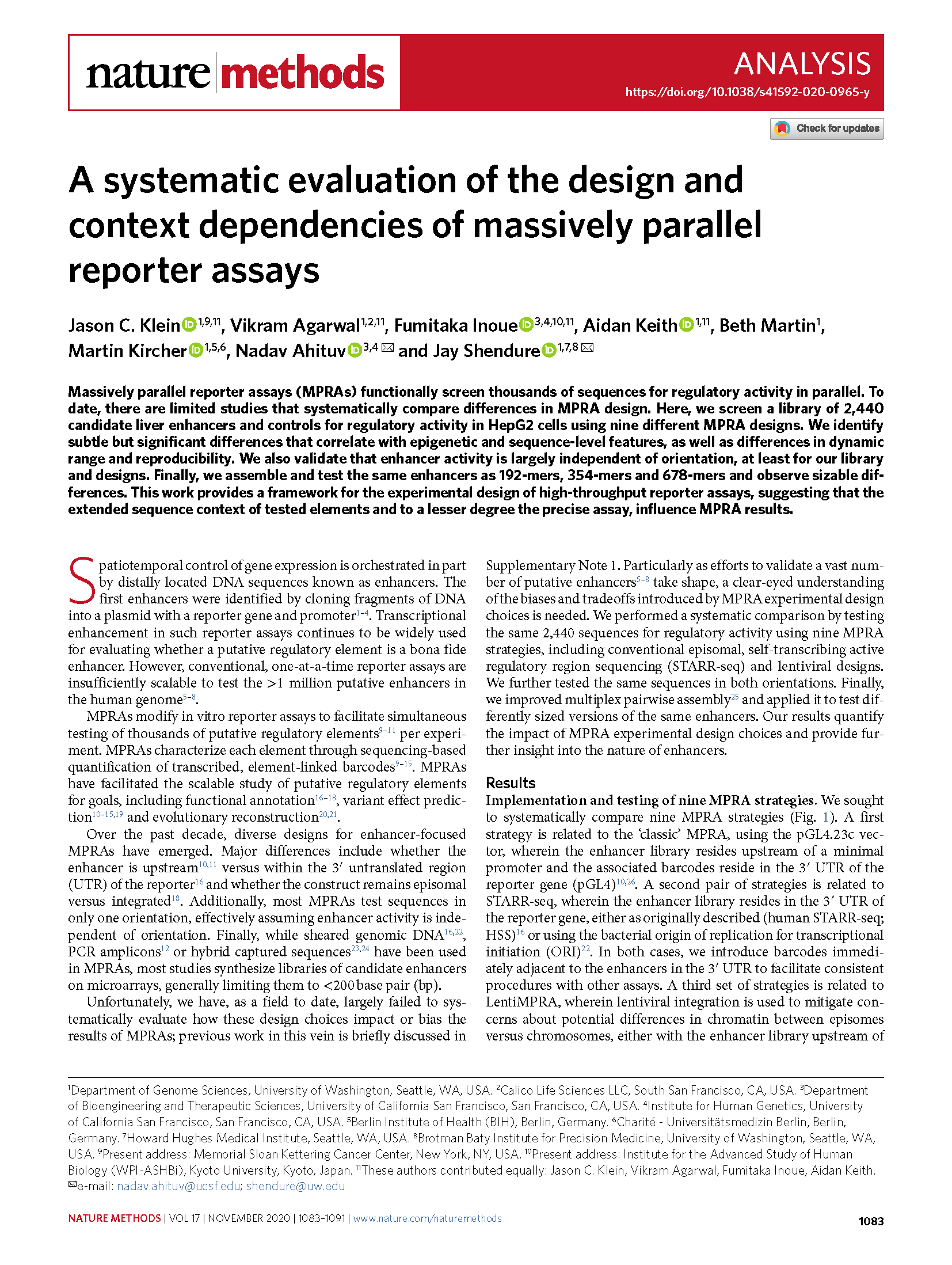
(opens in new tab) Klein, Agarwal, Inoue, Keith et al. A systematic evaluation of the design and context dependencies of massively parallel reporter assays
 (opens in new tab)
(opens in new tab) Bedford, Greninger,Roychoudhury, Starita, Famulare et al. Cryptic transmission of SARS-CoV-2 in Washington state
 (opens in new tab)
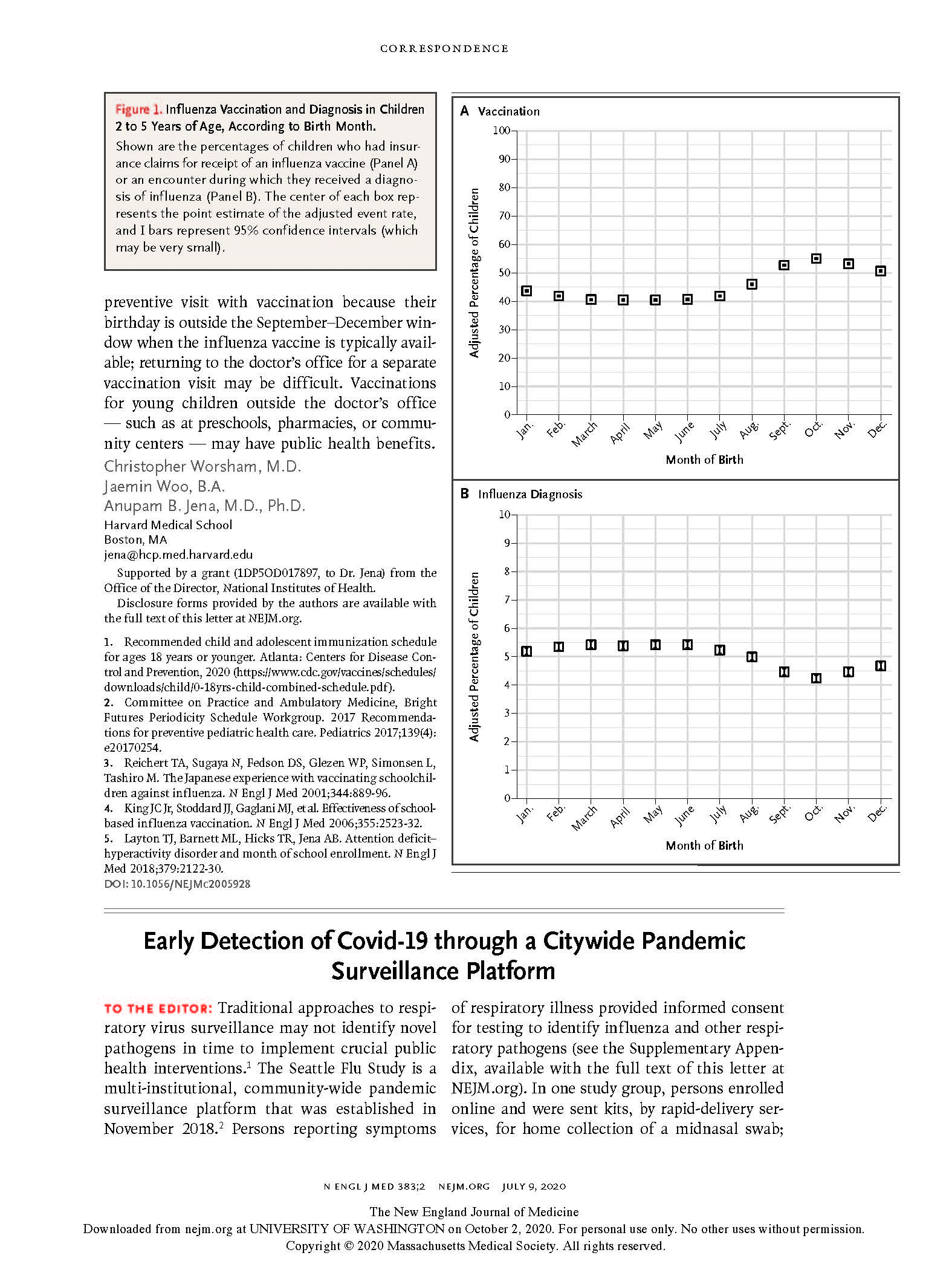
(opens in new tab) Chu et al. Early Detection of Covid-19 through a Citywide Pandemic Surveillance Platform
 (opens in new tab)
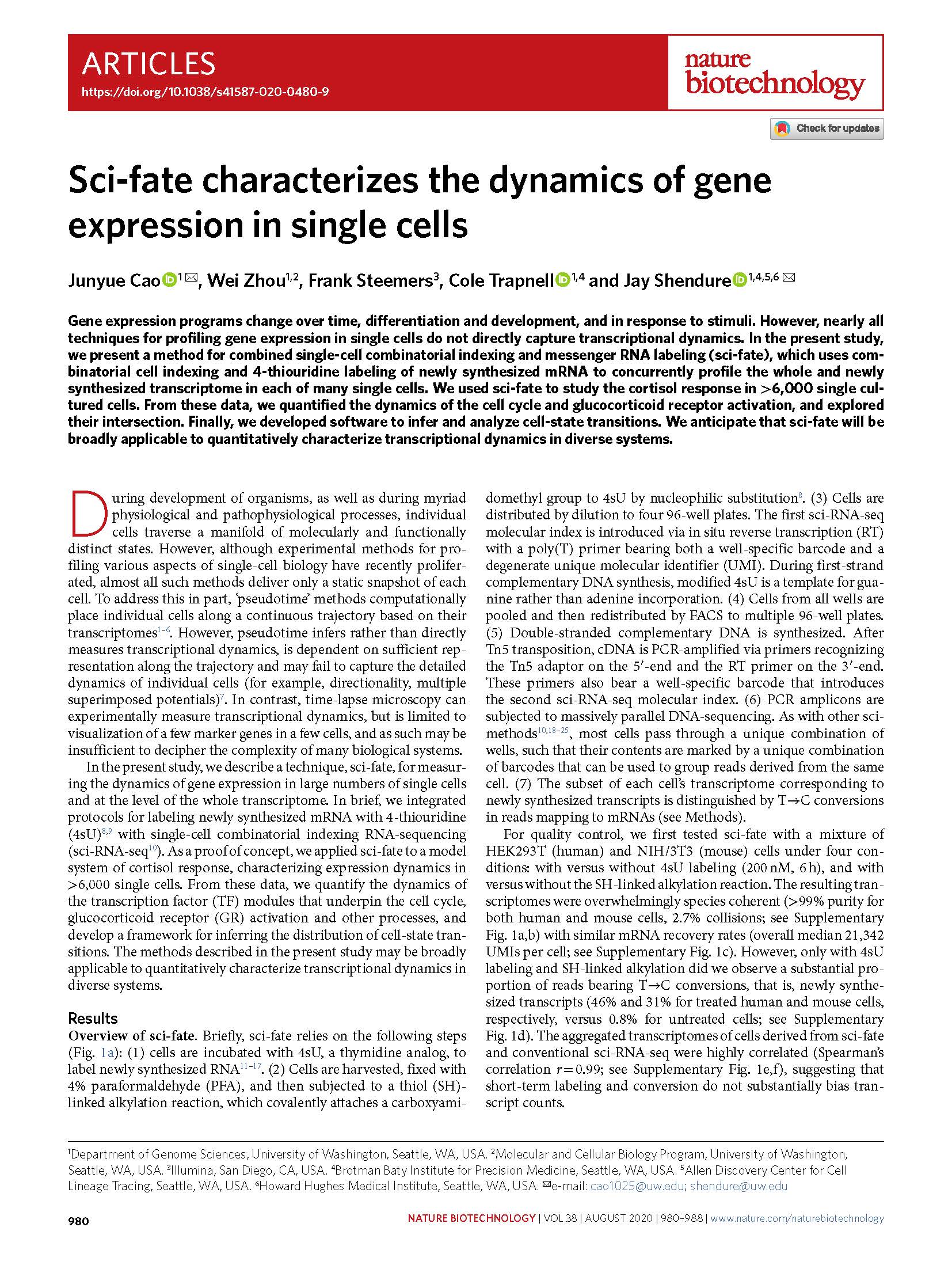
(opens in new tab) Cao et al. Sci-fate characterizes the dynamics of gene expression in single cells
 (opens in new tab)
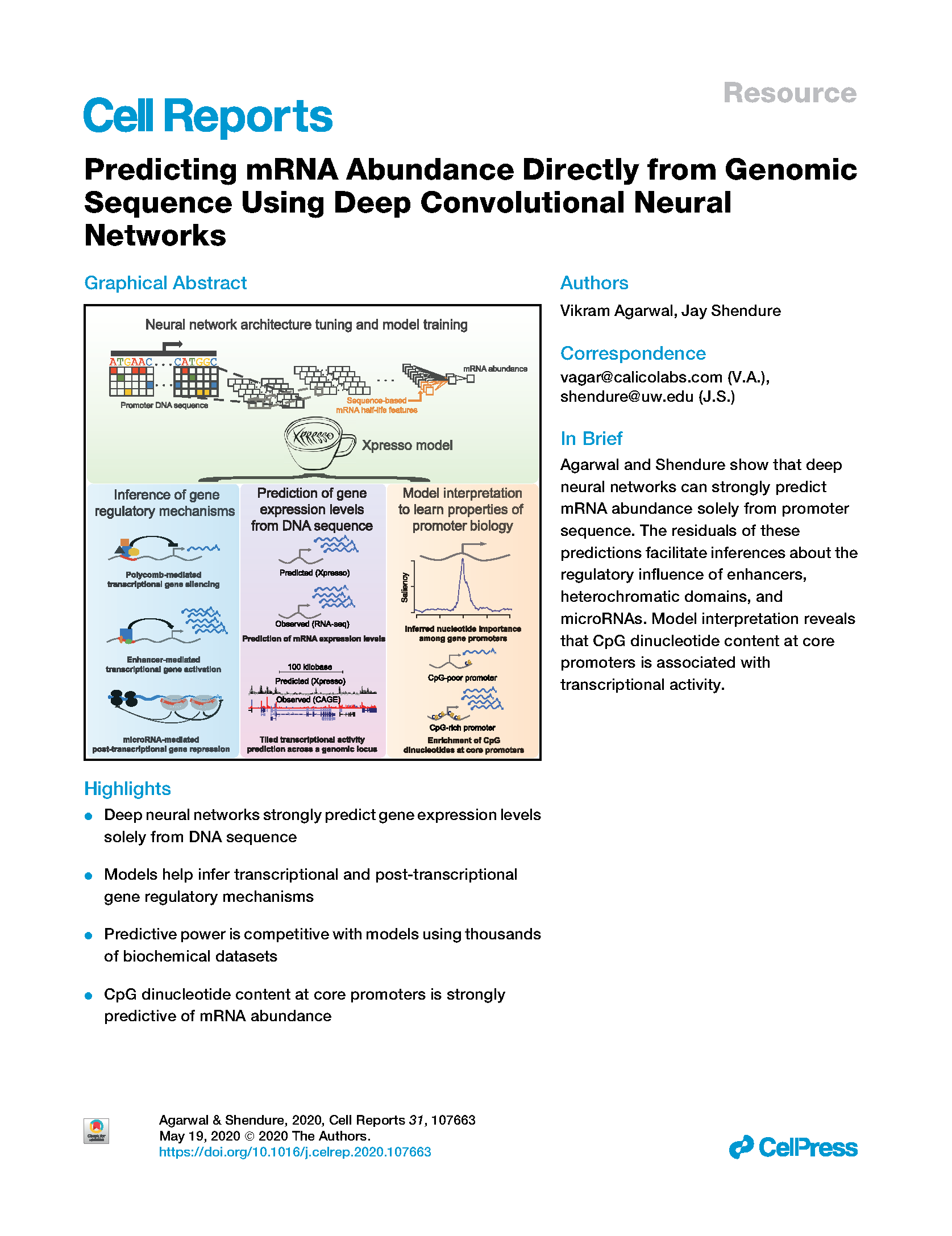
(opens in new tab) Agarwal, Shendure. Predicting mRNA abundance directly from genomic sequence using deep convolutional neural networks
 (opens in new tab)
(opens in new tab) Srivatsan, McFaline-Figueroa, Ramani et al. Massively multiplex chemical transcriptomics at single-cell resolution
2019
 (opens in new tab)
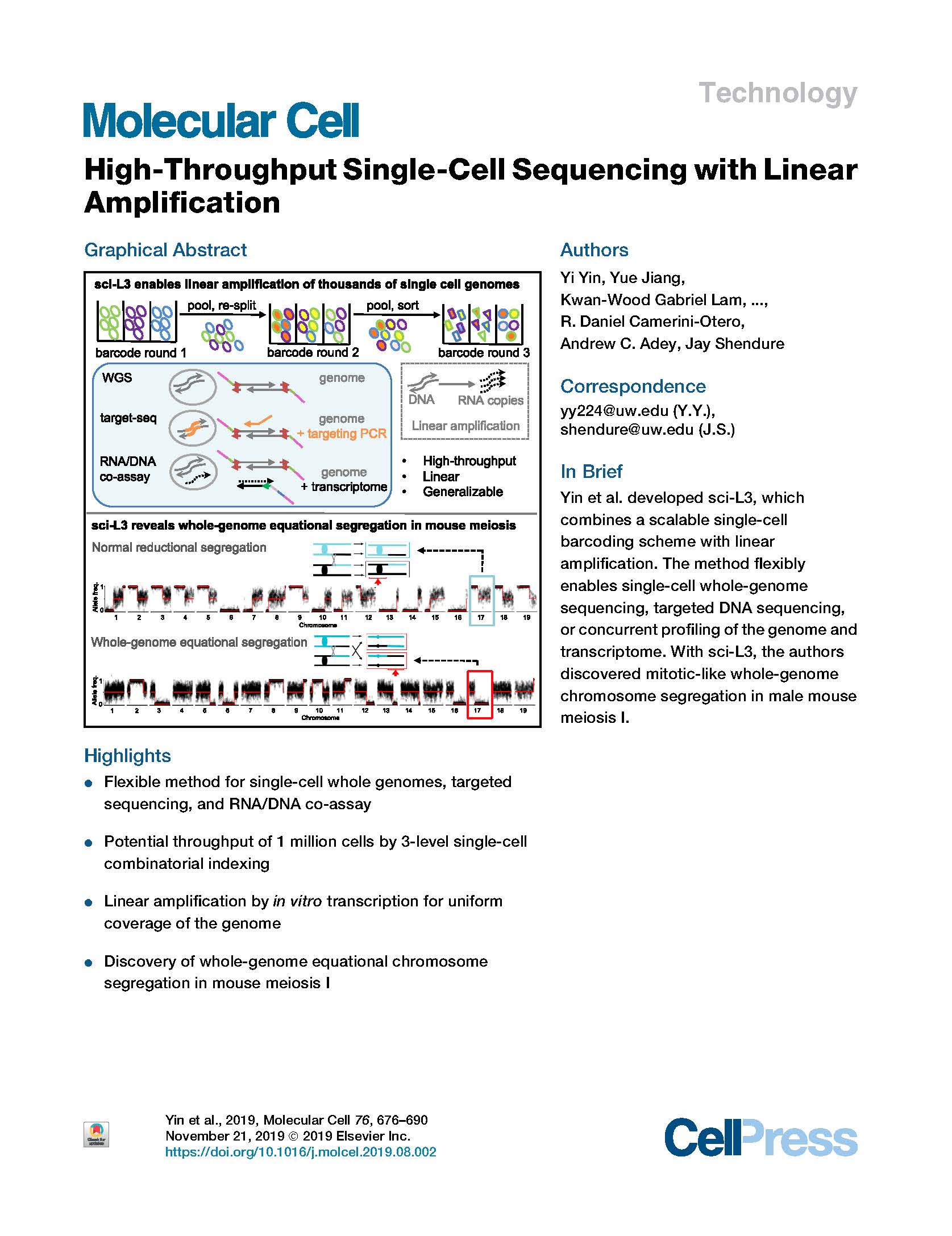
(opens in new tab) Yin et al. High-Throughput Single-Cell Sequencing with Linear Amplification
 (opens in new tab)
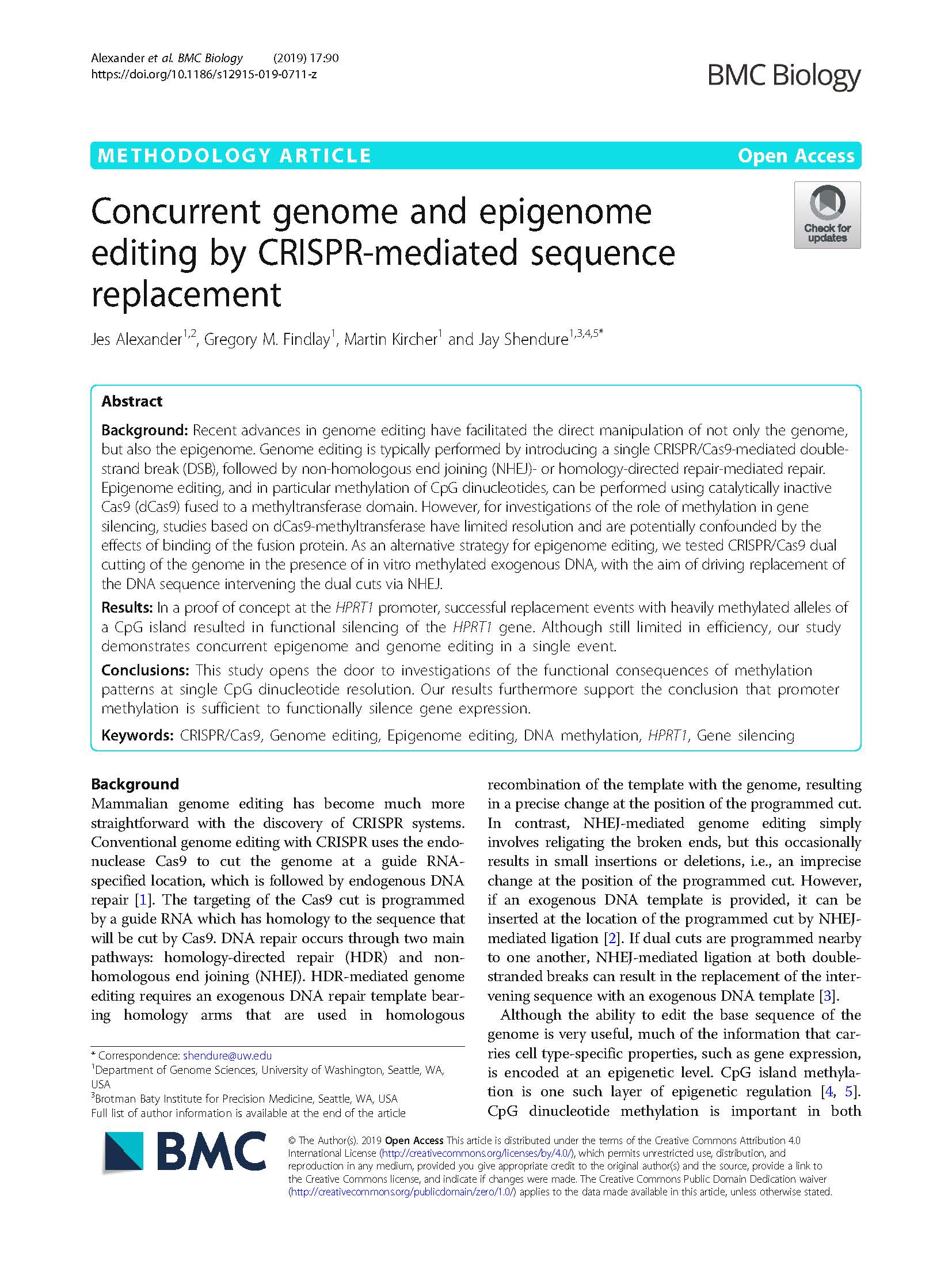
(opens in new tab) Alexander et al. Concurrent genome and epigenome editing by CRISPR-mediated sequence replacement
 (opens in new tab)
(opens in new tab) Pliner et al. Supervised classification enables rapid annotation of cell atlases
 (opens in new tab)
(opens in new tab) Kircher, Xiong, Martin, Schubach et al. Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution
 (opens in new tab)
(opens in new tab) Chen, McKenna, Schrieber et al. Massively parallel profiling and predictive modeling of the outcomes of CRISPR/Cas9-mediated double-strand break repair
 (opens in new tab)
(opens in new tab) Klein, Keith et al. Functional testing of thousands of osteoarthritis-associated variants for regulatory activity
 (opens in new tab)
(opens in new tab) Kim et al. A combination of transcription factors mediates inducible interchromosomal contacts
 (opens in new tab)
(opens in new tab) Ramani et al. High Sensitivity Profiling of Chromatin Structure by MNase-SSP
 (opens in new tab)
(opens in new tab) Cao, Spielmann et al. The single-cell transcriptional landscape of mammalian organogenesis
 (opens in new tab)
(opens in new tab) Gasperini et al. A Genome-wide Framework for Mapping Gene Regulation via Cellular Genetic Screens
 (opens in new tab)
(opens in new tab) Rentzsch et al. CADD: predicting the deleteriousness of variants throughout the human genome
2018
 (opens in new tab)
(opens in new tab) Starita et al. A Multiplex Homology-Directed DNA Repair Assay Reveals the Impact of More Than 1,000 BRCA1 Missense Substitution Variants on Protein Function
 (opens in new tab)
(opens in new tab) Findlay et al. Accurate classification of BRCA1 variants with saturation genome editing
 (opens in new tab)
(opens in new tab) Cao et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells
 (opens in new tab)
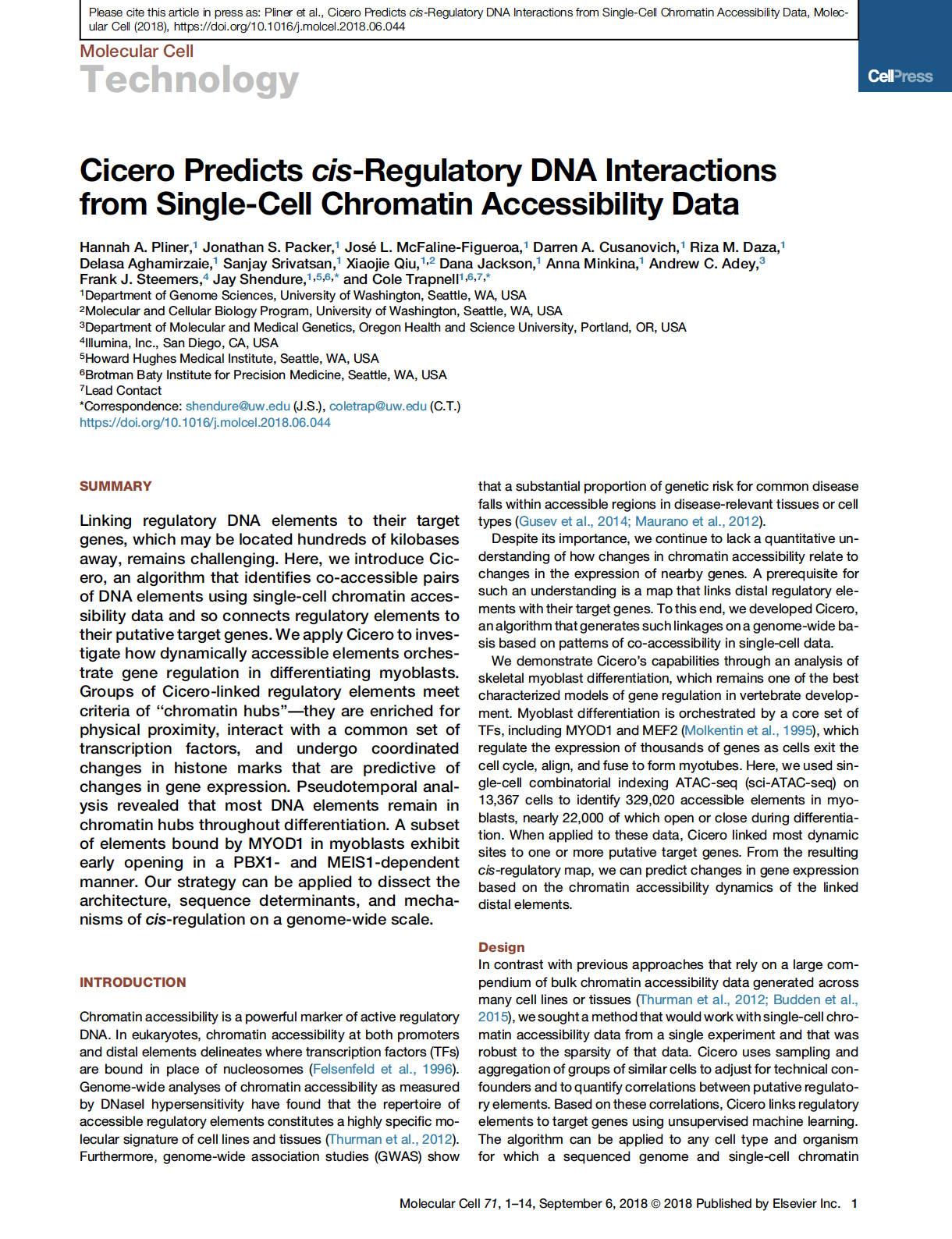
(opens in new tab) Pliner et al. Cicero Predicts cis-Regulatory DNA Interactions from Single-Cell Chromatin Accessibility Data
 (opens in new tab)
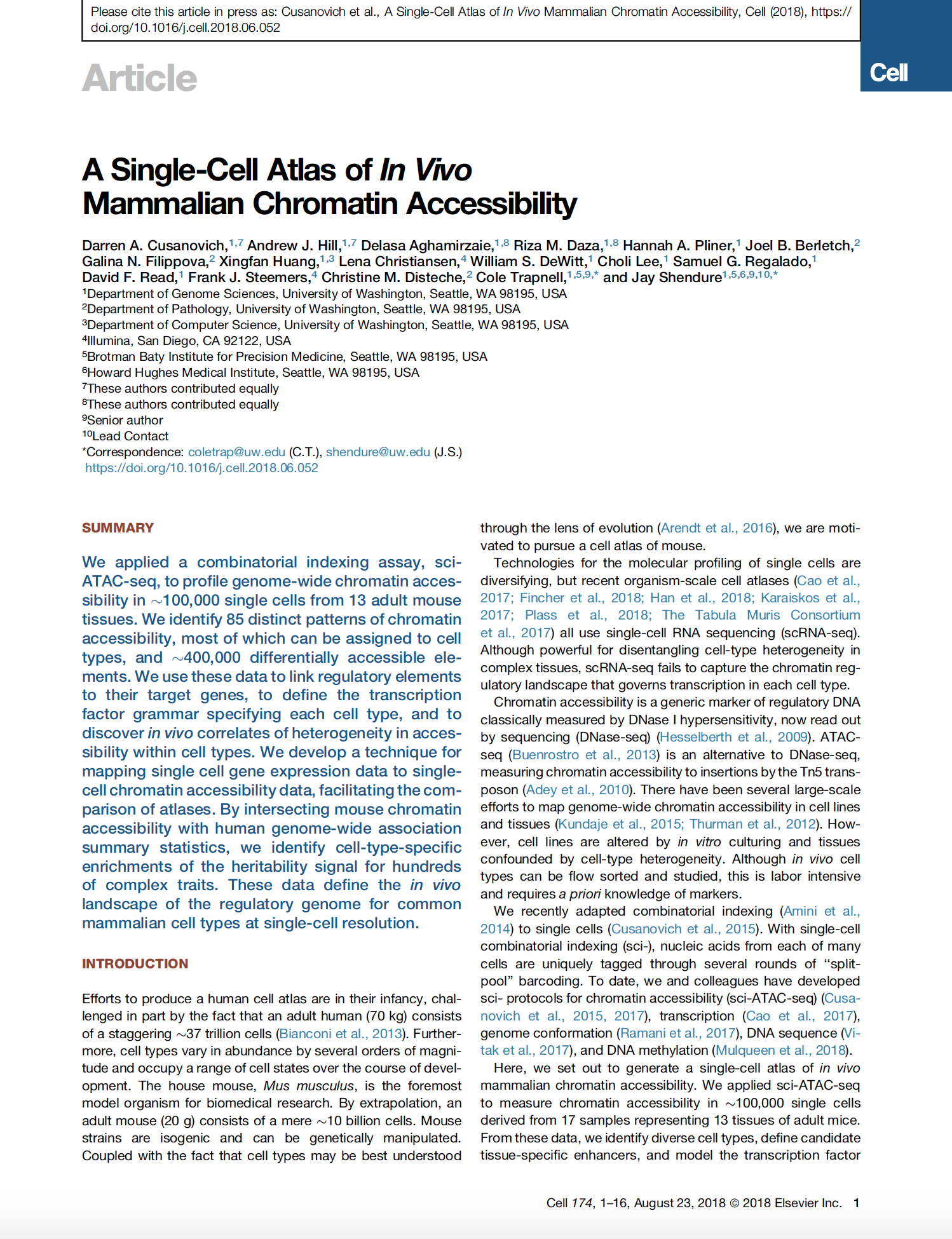
(opens in new tab) Cusanovich, Hill et al. A Single-Cell Atlas of In Vivo Mammalian Chromatin Accessibility
 (opens in new tab)
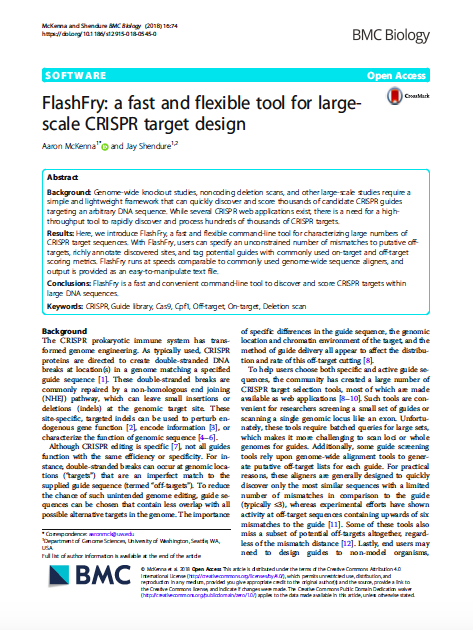
(opens in new tab) McKenna & Shendure. FlashFry: a fast and flexible tool for large-scale CRISPR target design
 (opens in new tab)
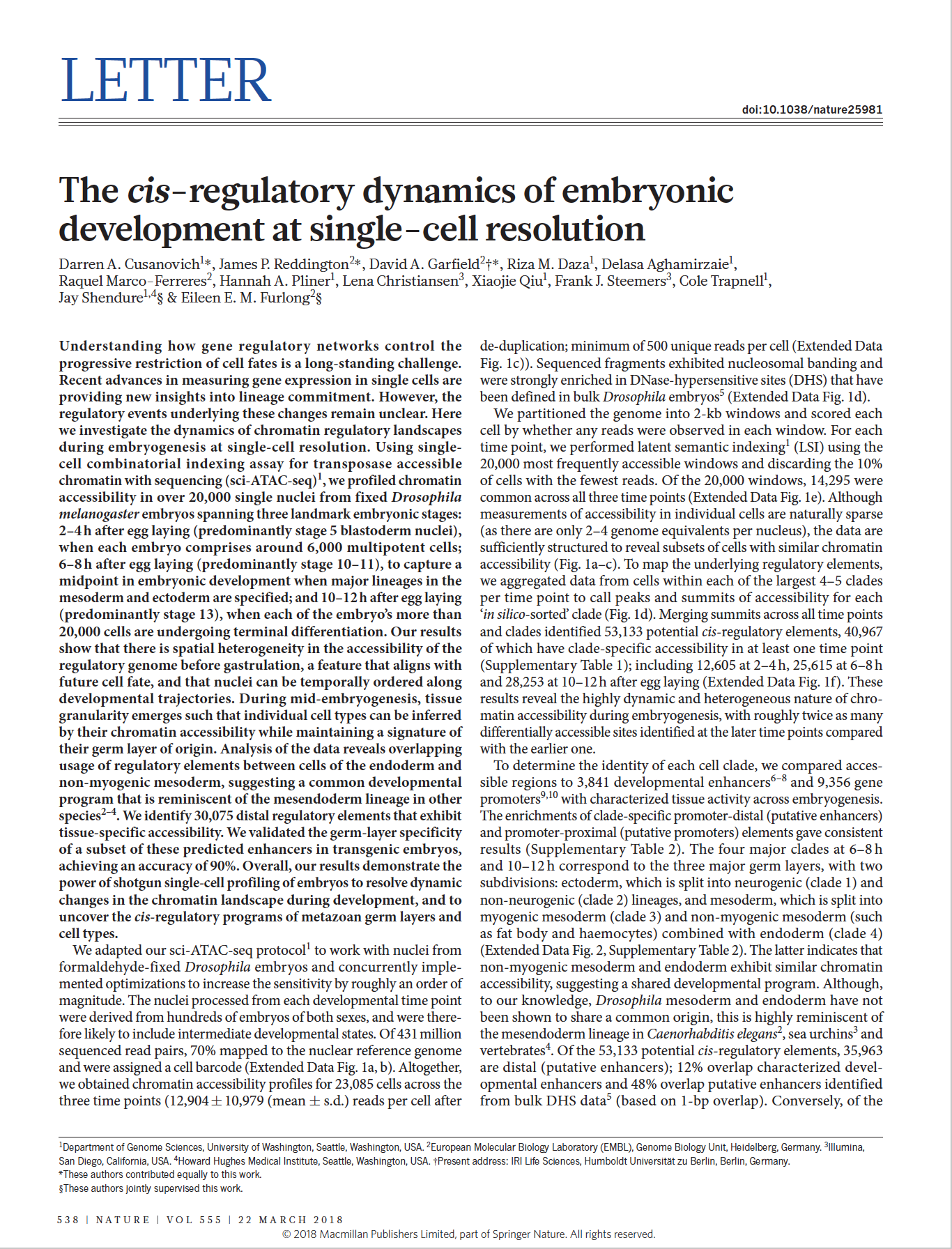
(opens in new tab) Cusanovich, Reddington, Garfield et al. The cis-regulatory dynamics of embryonic development at single-cell resolution
 (opens in new tab)
(opens in new tab) Klein et al. Functional characterization of enhancer evolution in the primate lineage
 (opens in new tab)
(opens in new tab) Hill, McFaline-Figueroa et al. On the design of CRISPR-based single-cell molecular screens
 (opens in new tab)
(opens in new tab) Matreyek, Starita et al. Multiplex assessment of protein variant abundance by massively parallel sequencing
2017
 (opens in new tab)
(opens in new tab) Cao, Packer et al. Comprehensive single cell transcriptional profiling of a multicelluar organism by combinatorial indexing
 (opens in new tab)
(opens in new tab) Kim et al. The dynamic three-dimensional organization of the diploid yeast genome
 (opens in new tab)
(opens in new tab) Gasperini, Findlay et al. CRISPR/Cas9-Mediated Scanning for Regulatory Elements Required for HPRT1 Expression via Thousands of Large, Programmed Genomic Deletions
 (opens in new tab)
(opens in new tab) Neveling, Mensenkamp et al. BRCA Testing by Single-Molecule Molecular Inversion Probes
 (opens in new tab)
(opens in new tab) Inoue, Kircher et al. A systematic comparison reveals substantial differences in chromosomal versus episomal encoding of enhancer activity
 (opens in new tab)
(opens in new tab) Ramani et al. Massively multiplex single-cell Hi-C
2016
 (opens in new tab)
(opens in new tab) Hause et al. Classification and characterization of microsatellite instability across 18 cancer types
 (opens in new tab)
(opens in new tab) Ramani, Cusanovich et al. Mapping 3D genome architecture through in situ DNase Hi-C
 (opens in new tab)
(opens in new tab) Snyder, Kircher et al. Cell-free DNA Comprises an In Vivo Nucleosome Footprint that Informs Its Tissues-Of-Origin
 (opens in new tab)
(opens in new tab) Kumar, Coleman et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer
 (opens in new tab)
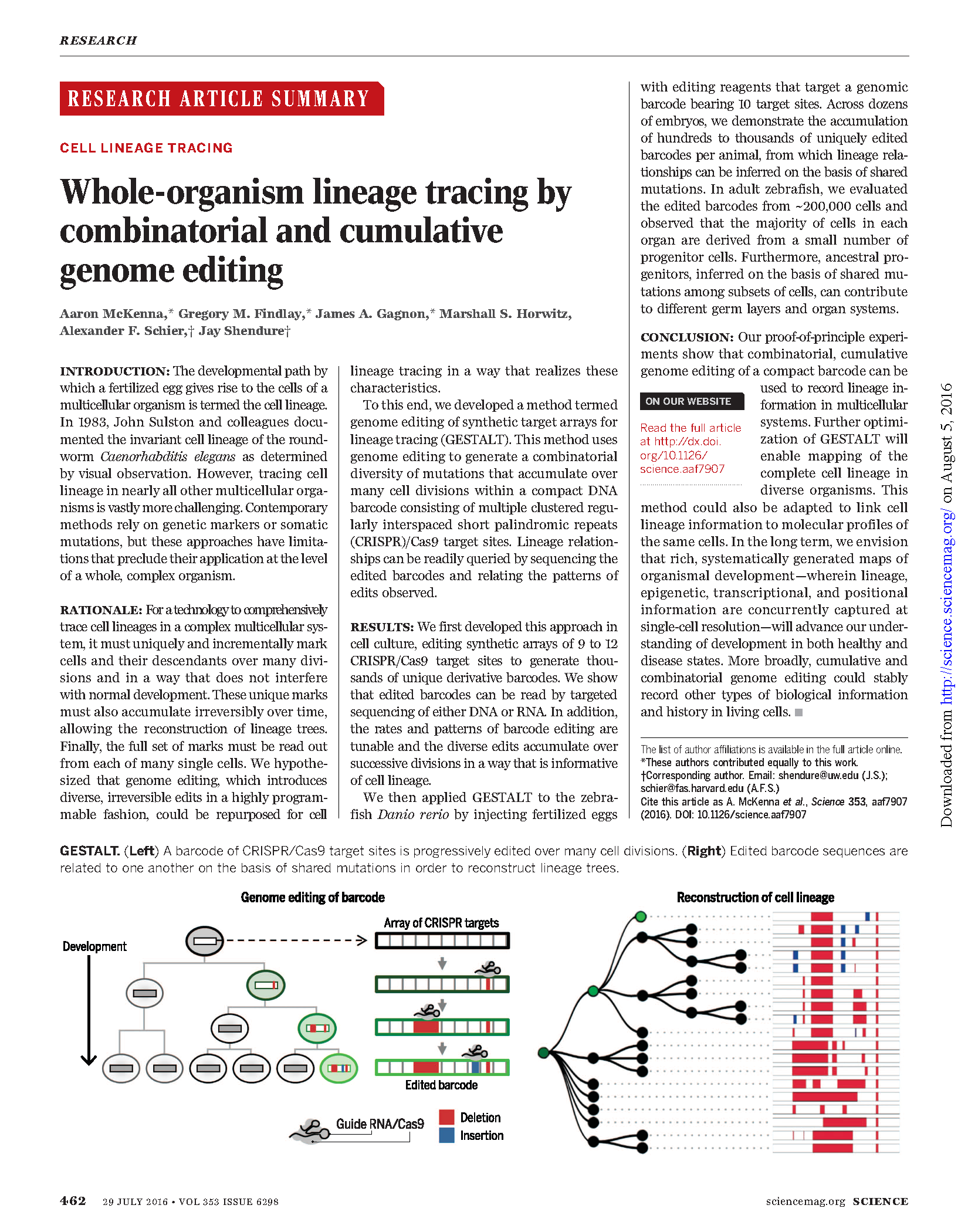
(opens in new tab) McKenna, Findlay, Gagnon et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing
 (opens in new tab)
(opens in new tab) Underhill et al. Fragment Length of Circulating Tumor DNA
2015
 (opens in new tab)
(opens in new tab) Ramani et al. High-throughput determination of RNA structure by proximity ligation
 (opens in new tab)
(opens in new tab) Fairfield et al. Exome sequencing reveals pathogenic mutations in 91 strains of mice with Mendelian disorders
 (opens in new tab)
(opens in new tab) Starita et al. Massively Parallel Functional Analysis of BRCA1 RING Domain Variants
 (opens in new tab)
(opens in new tab) Klein et al. Multiplex pairwise assembly of array-derived DNA oligonucleotides
 (opens in new tab)
(opens in new tab) Roach et al. A Year of Infection in the Intensive Care Unit: Prospective Whole Genome Sequencing of Bacterial Clinical Isolates Reveals Cryptic Transmissions and Novel Microbiota
 (opens in new tab)
(opens in new tab) Kitzman, Starita et al. Massively parallel single-amino-acid mutagenesis
 (opens in new tab)
(opens in new tab) Cusanovich et al. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing
 (opens in new tab)
(opens in new tab) Kumar, Ryan et al. Whole genome prediction for preimplantation genetic diagnosis
 (opens in new tab)
(opens in new tab) Snyder, Simmons et al. Copy-Number Variation and False Positive Prenatal Aneuploidy Screening Results
2014
 (opens in new tab)
(opens in new tab) O'Roak, Stessman et al. Recurrent de novo mutations implicate novel genes underlying simplex autism risk
 (opens in new tab)
(opens in new tab) Kumar et al. Deep sequencing of multiple regions of glial tumors reveals spatial heterogeneity for mutations in clinically relevant genes
 (opens in new tab)
(opens in new tab) Salipante, Roach et al. Large-scale genomic sequencing of extraintestinal pathogenic Escherichia coli strains
 (opens in new tab)
(opens in new tab) Findlay, Boyle et al. Saturation editing of genomic regions by multiplex homology-directed repair
 (opens in new tab)
(opens in new tab) Adey et al. In vitro, long-range sequence information for de novo genome assembly via transposase contiguity
 (opens in new tab)
(opens in new tab) Burton, Liachko et al. Species-Level Deconvolution of Metagenome Assemblies with Hi-C–Based Contact Probability Maps. G3 (2014)
 (opens in new tab)
(opens in new tab) Iossifov, O'Roak, Sanders, Ronemus et al. The contribution of de novo coding mutations to autism spectrum disorder
 (opens in new tab)
(opens in new tab) Schwartz, Roach et al. Primate evolution of the recombination regulator PRDM9
 (opens in new tab)
(opens in new tab) Boyle et al. MIPgen: optimized modeling and design of molecular inversion probes for targeted resequencing
 (opens in new tab)
(opens in new tab) Kircher, Witten et al. A general framework for estimating the relative pathogenicity of human genetic variants
2013
 (opens in new tab)
(opens in new tab) Burton et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions
 (opens in new tab)
(opens in new tab) Hiatt et al. Single molecule molecular inversion probes for targeted, high-accuracy detection of low-frequency variation
 (opens in new tab)
(opens in new tab) Adey, Burton, Kitzman et al. The haplotype-resolved genome and epigenome of the aneuploid HeLa cancer cell line
 (opens in new tab)
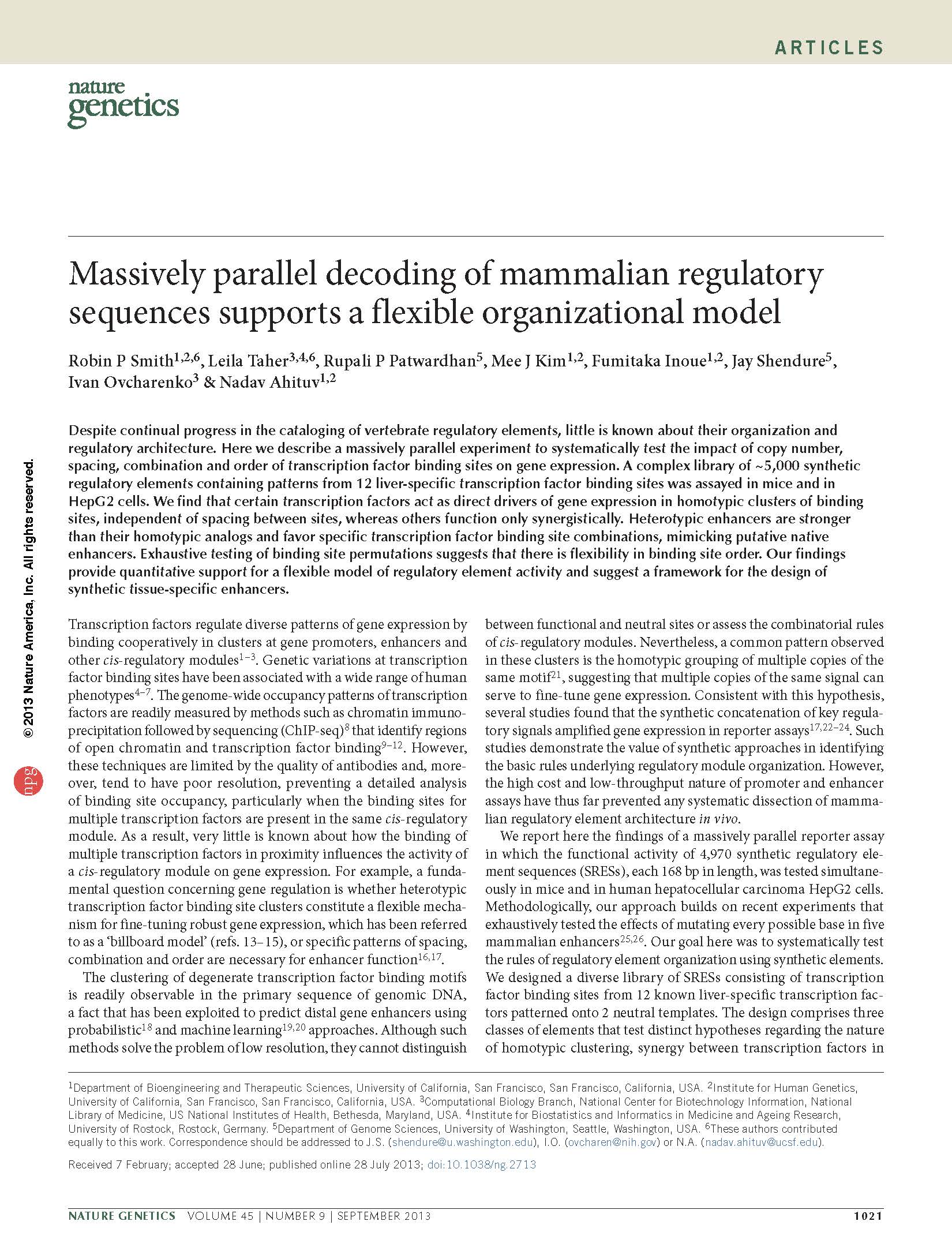
(opens in new tab) Smith, Taher L et al. Massively parallel decoding of mammalian regulatory sequences supports a flexible organizational model
2012
 (opens in new tab)
(opens in new tab) O'Roak et al. Multiplex Targeted Sequencing Identifies Recurrently Mutated Genes in Autism Spectrum Disorders
 (opens in new tab)
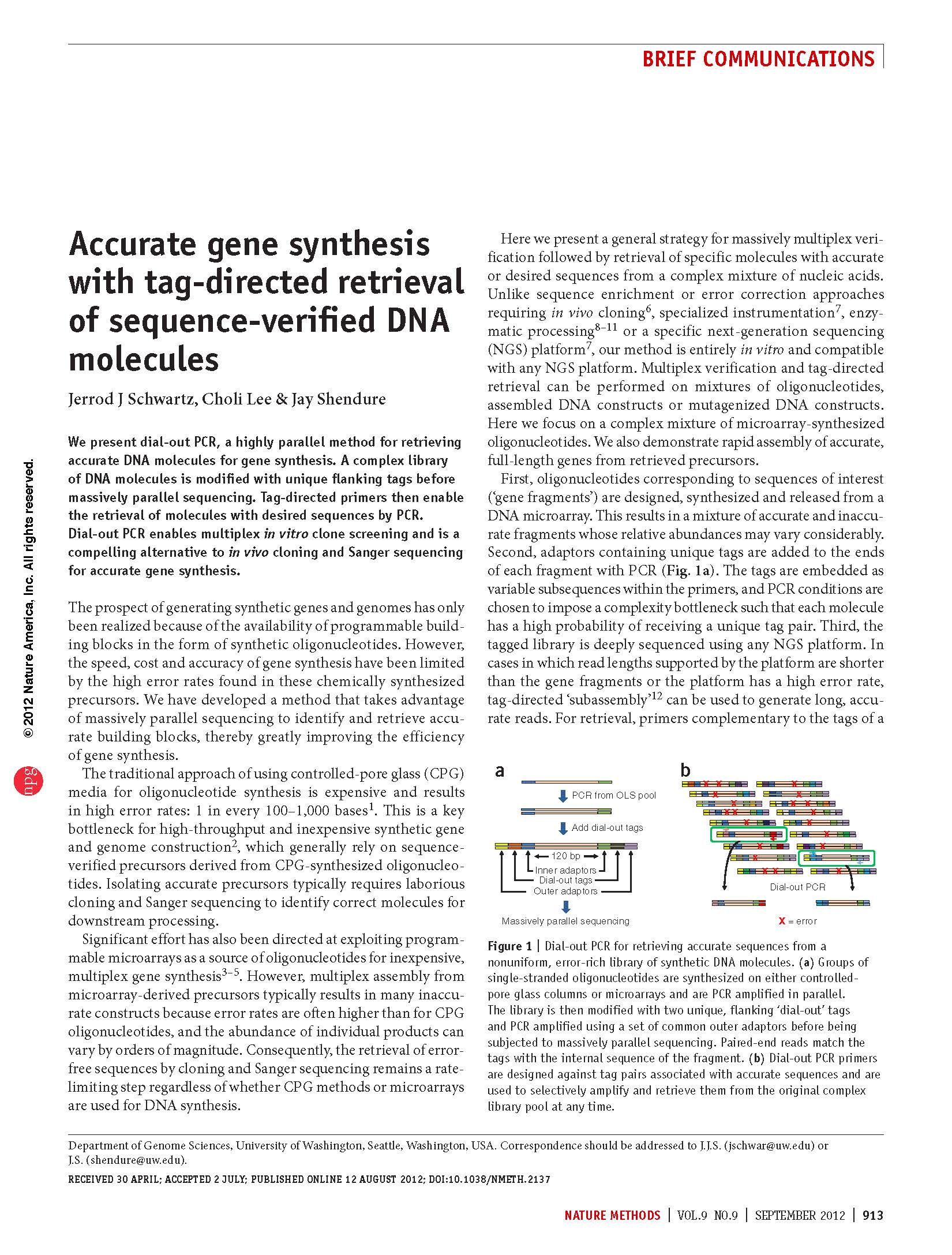
(opens in new tab) Schwartz et al. Accurate gene synthesis with tag-directed retrieval of sequence-verified DNA molecules
 (opens in new tab)
(opens in new tab) Schwartz et al. Capturing native long-range contiguity by in situ library construction and optical sequencing
 (opens in new tab)
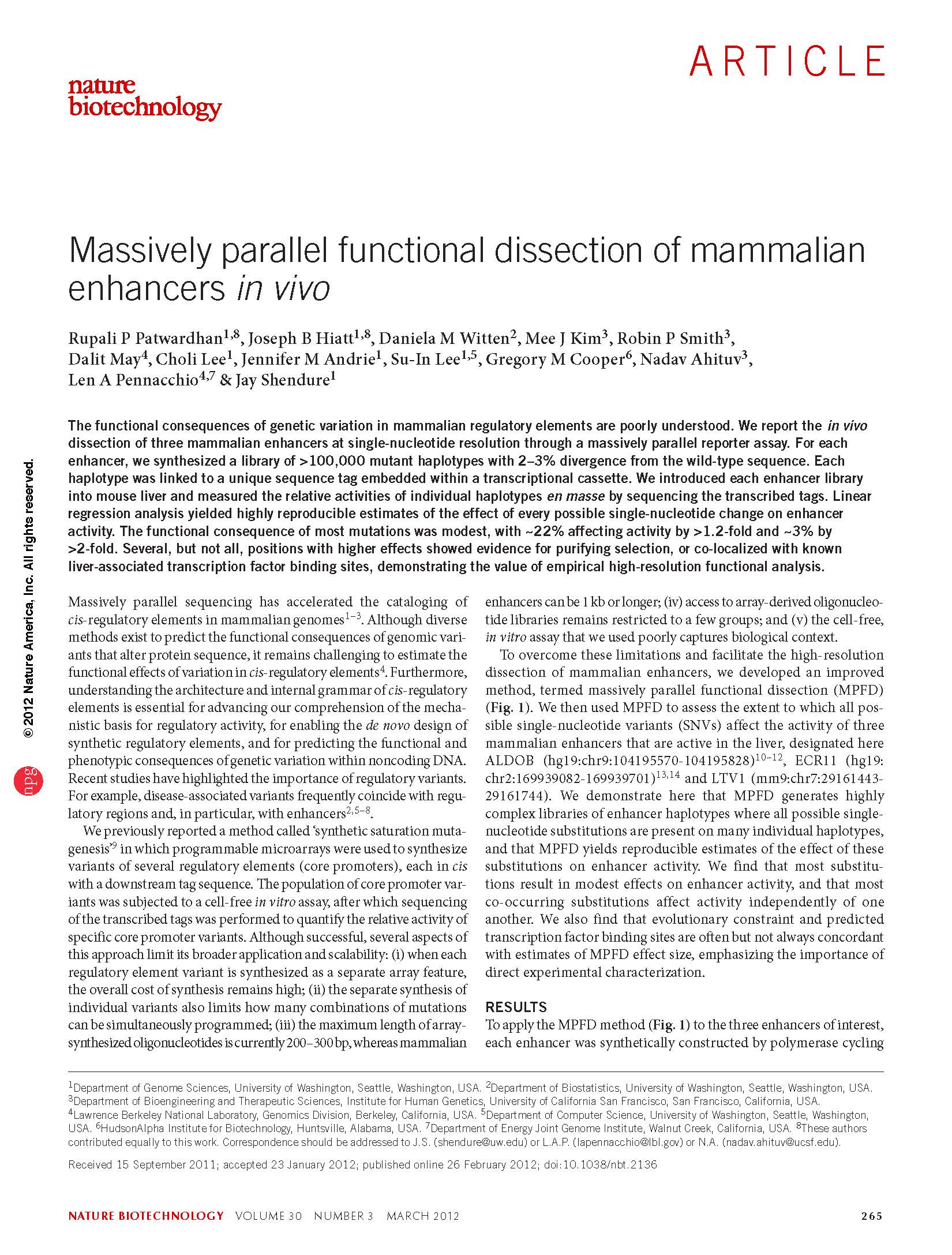
(opens in new tab) Patwardhan, Hiatt et al. Massively parallel functional dissection of mammalian enhancers in vivo
 (opens in new tab)
(opens in new tab) Kitzman et al. Noninvasive whole-genome sequencing of a human fetus
 (opens in new tab)
(opens in new tab) O'Roak et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations
 (opens in new tab)
(opens in new tab) Adey, Shendure. Ultra-low-input, tagmentation-based whole-genome bisulfite sequencing
2011
 (opens in new tab)
(opens in new tab) Kumar et al. Exome sequencing identifies a spectrum of mutation frequencies in advanced and lethal prostate cancers
 (opens in new tab)
(opens in new tab) George et al. Trans genomic capture and sequencing of primate exomes reveals new targets of positive selection
 (opens in new tab)
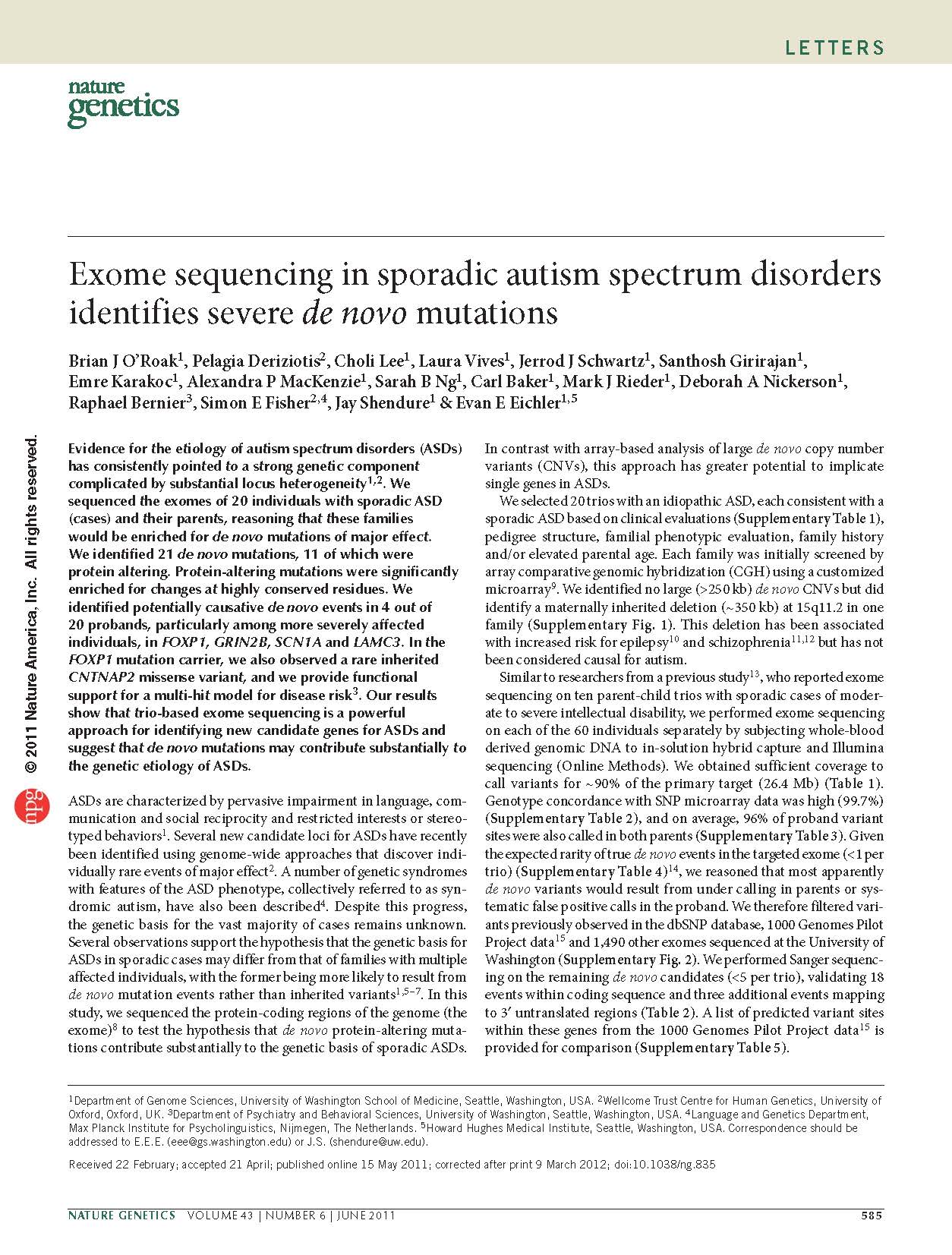
(opens in new tab) O'Roak et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations
 (opens in new tab)
(opens in new tab) Kitzman et al. Haplotype-resolved genome sequencing of a Gujarati Indian individual
2010
 (opens in new tab)
(opens in new tab) Ng, Bigham et al. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome
 (opens in new tab)
(opens in new tab) Hiatt, Patwardhan et al. Parallel, tag-directed assembly of locally derived short sequence reads
 (opens in new tab)
(opens in new tab) Adey, Morrison, Asan, Xun et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high-density in vitro transposition
 (opens in new tab)
(opens in new tab) Ng, Buckingham et al. Exome sequencing identifies the cause of a mendelian disorder
2009
 (opens in new tab)
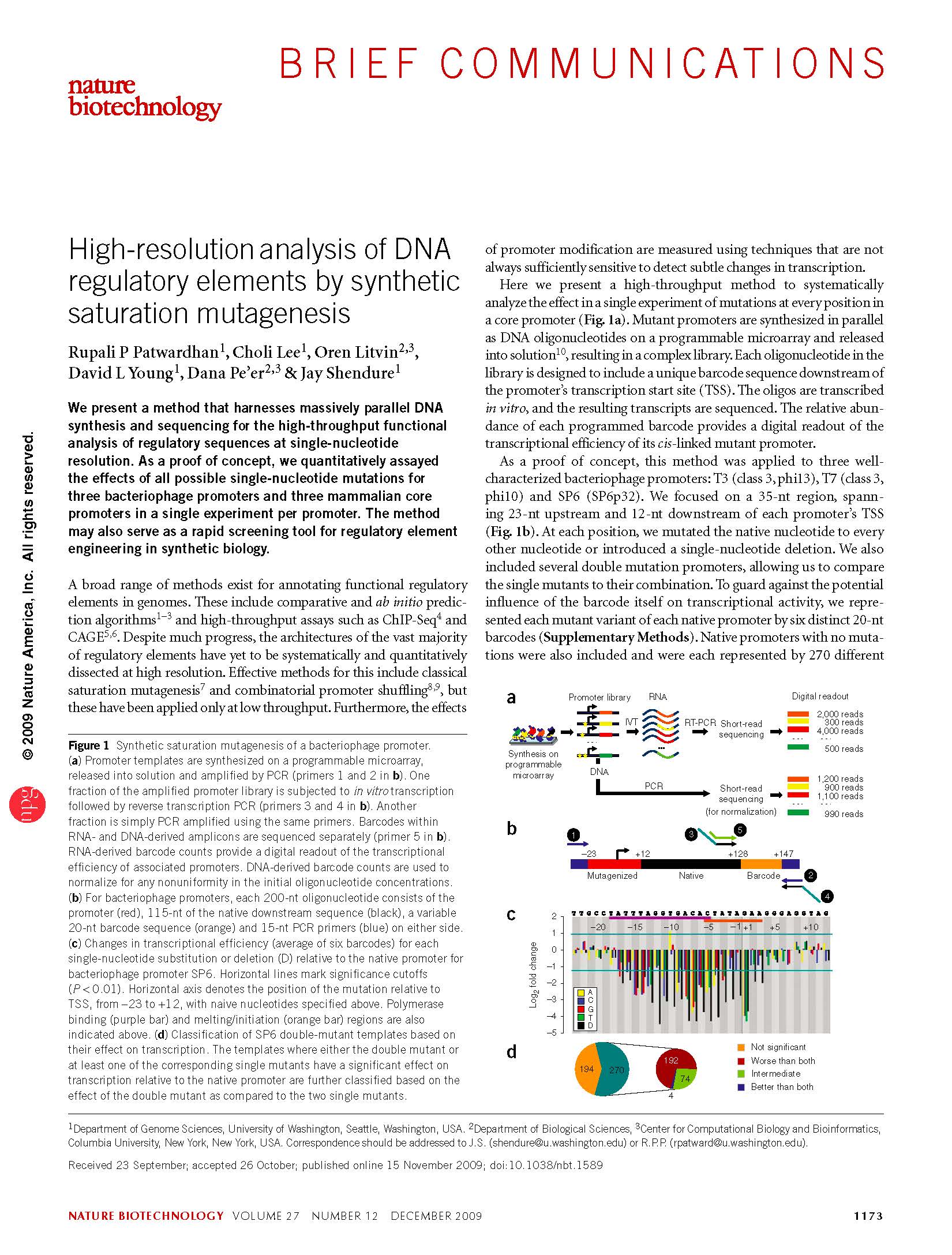
(opens in new tab) Patwardhan et al. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis
 (opens in new tab)
(opens in new tab) Ng et al. Targeted capture and massively parallel sequencing of 12 human exomes
 (opens in new tab)
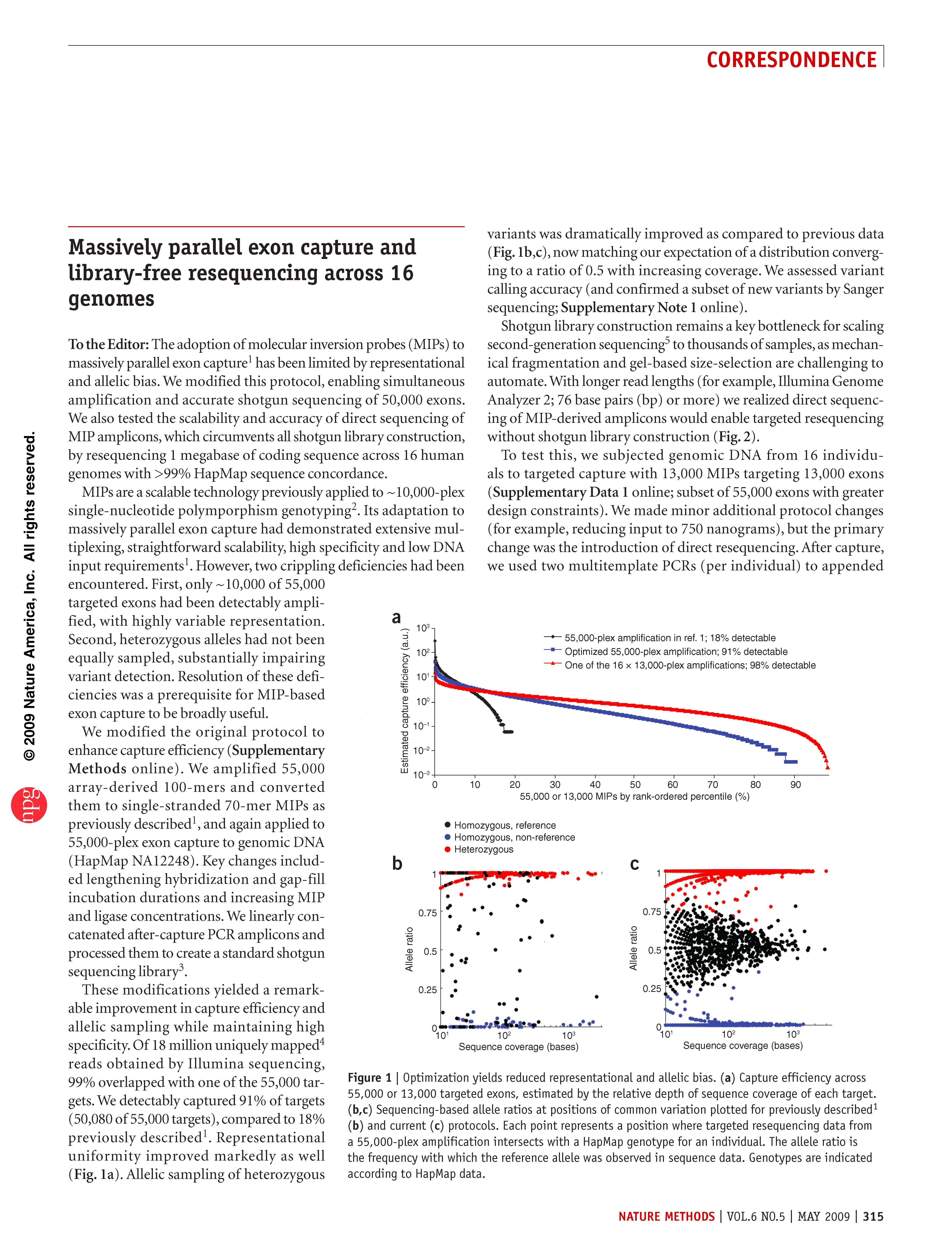
(opens in new tab) Turner et al. Massively parallel exon capture and library-free resequencing across 16 genomes
 (opens in new tab)
(opens in new tab) Vasta V et al. Next generation sequence analysis for mitochondrial disorders