Massively Parallel Functional Genomics
(selected publications)
2025
 (opens in new tab)
(opens in new tab) Suiter et al. Combinatorial mapping of E3 ubiquitin ligases to their target substrates.
Molecular Cell (2025)
 (opens in new tab)
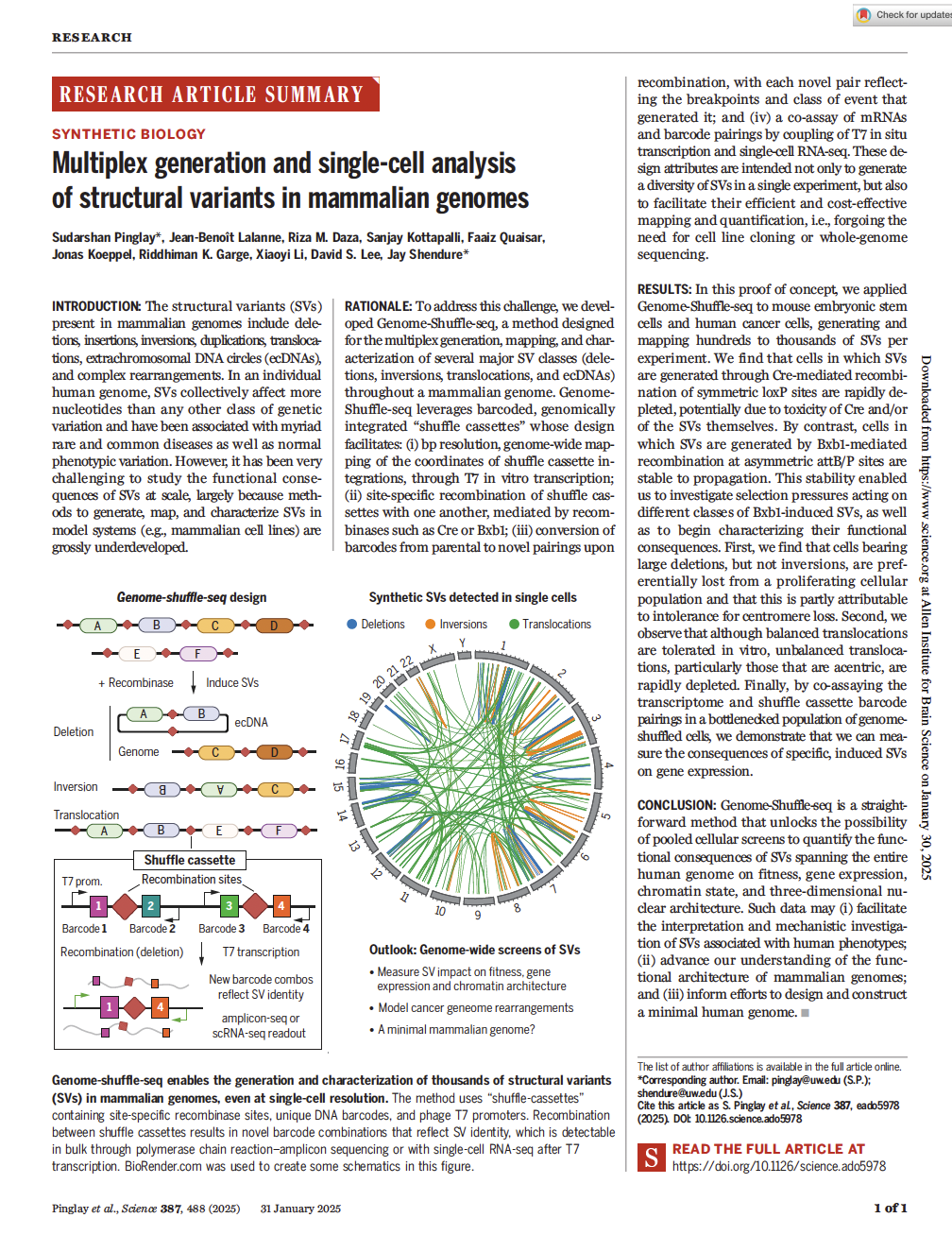
(opens in new tab) Pinglay et al. Multiplex generation and single-cell analysis of structural variants in mammalian genomes.
Science (2025)
 (opens in new tab)
(opens in new tab) Agarwal, Inoue et al. Massively parallel characterization of transcriptional regulatory elements.
Nature (2025)
2024
 (opens in new tab)
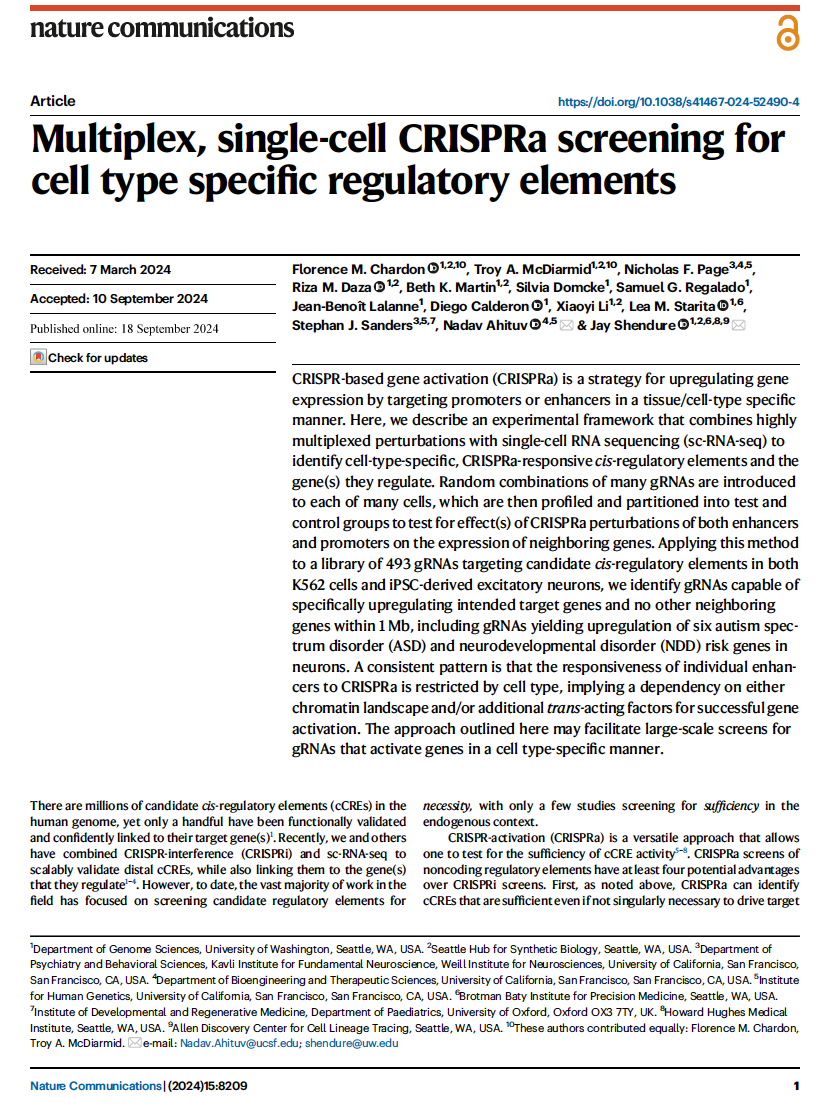
(opens in new tab) Chardon, McDiarmid et al. Multiplex, single-cell CRISPRa screening for cell type specific regulatory elements.
Nature Communications (2024)
 (opens in new tab)
(opens in new tab) Lalanne, Regalado et al. Multiplex profiling of developmental cis-regulatory elements with quantitative single-cell expression reporters.
Nature Methods (2024)
2020
 (opens in new tab)
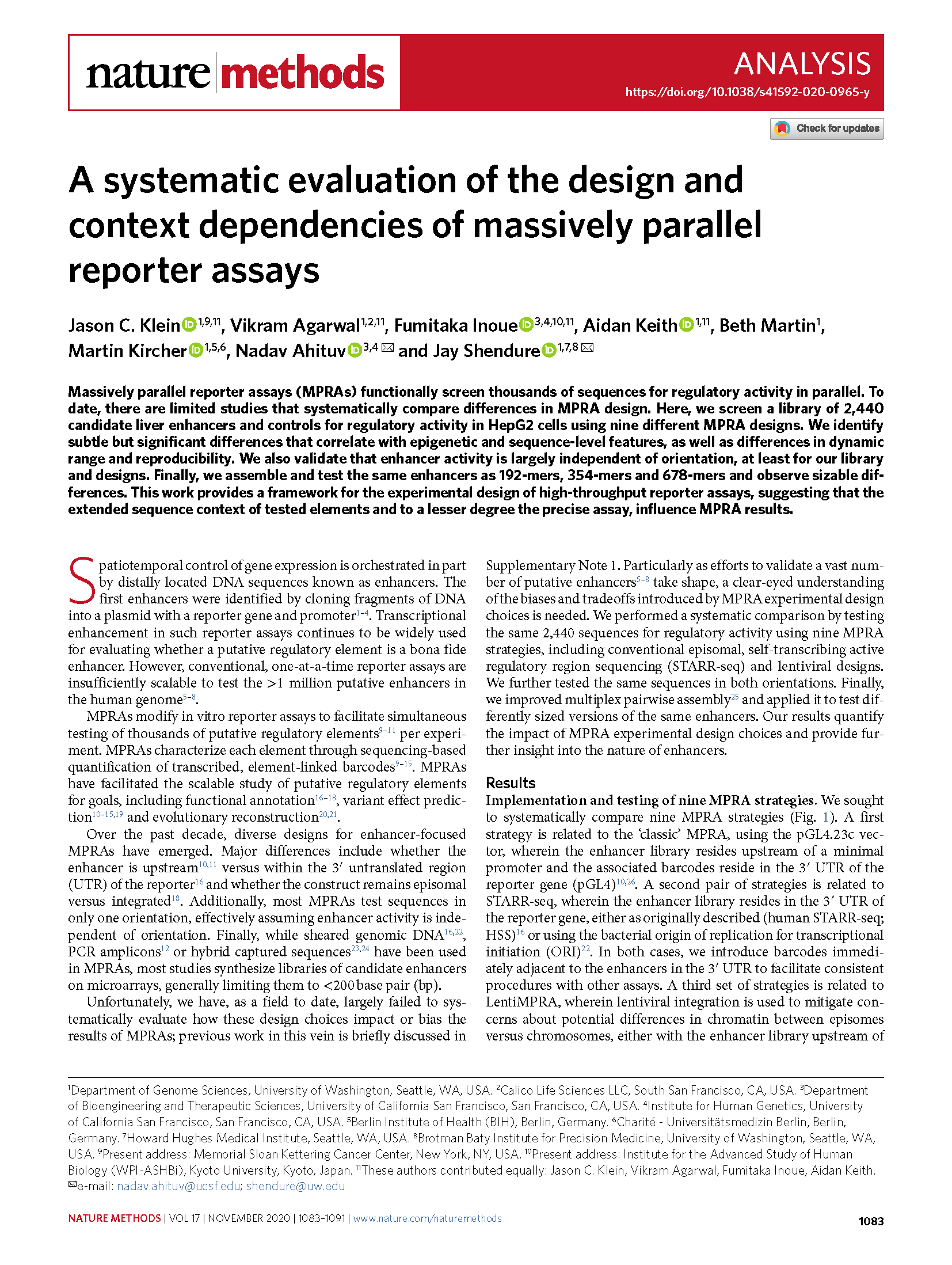
(opens in new tab) Klein, Agarwal, Inoue, Keith et al. A systematic evaluation of the design and context dependencies of massively parallel reporter assays.
Nature Methods (2020)
 (opens in new tab)
(opens in new tab) Gordon, Inoue, Martin, Schubach et al. lentiMPRA and MPRAflow for high-throughput functional characterization of gene regulatory elements.
Nature Protocols (2020)
PMID: 32641802 (opens in new tab) | PDF (opens in new tab)
(with Ahituv and Kircher Labs)
 (opens in new tab)
(opens in new tab) Gasperini, Tome, Shendure. Towards a comprehensive catalogue of validated and target-linked human enhancers.
Nature Reviews Genetics (2020)
 (opens in new tab)
(opens in new tab) Srivatsan, McFaline-Figueroa, Ramani et al. Massively multiplex chemical transcriptomics at single-cell resolution.
Science (2020)
PMID: 31806696 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
2019
 (opens in new tab)
(opens in new tab) Kircher, Xiong, Martin, Schubach et al. Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution.
Nature Communications (2019)
PMID: 31395865 (opens in new tab) | PDF (opens in new tab)
(with Ahituv and Kircher Labs)
 (opens in new tab)
(opens in new tab) Chen, McKenna, Schreiber et al. Massively parallel profiling and predictive modeling of the outcomes of CRISPR/Cas9-mediated double-strand break repair.
Nucleic Acids Research (2019)
 (opens in new tab)
(opens in new tab) Klein, Keith et al. Functional testing of thousands of osteoarthritis-associated variants for regulatory activity.
Nature Communications (2019)
 (opens in new tab)
(opens in new tab) Kim, Dunham, Shendure. A combination of transcription factors mediates inducible interchromosomal contacts.
eLife (2019)
 (opens in new tab)
(opens in new tab) Gasperini et al. A Genome-wide Framework for Mapping Gene Regulation via Cellular Genetic Screens.
Cell (2019)
2018
 (opens in new tab)
(opens in new tab) Findlay et al. Accurate classification of BRCA1 variants with saturation genome editing.
Nature (2018)
PMID: 30209399 (opens in new tab) | PDF (opens in new tab)
(with Starita Lab)
 (opens in new tab)
(opens in new tab) Klein et al. Functional characterization of enhancer evolution in the primate lineage.
Genome Biology (2018)
 (opens in new tab)
(opens in new tab) Starita et al. A Multiplex Homology-Directed DNA Repair Assay Reveals the Impact of More Than 1,000 BRCA1 Missense Substitution Variants on Protein Function.
AJHG (2018)
PMID: 30219179 (opens in new tab) | PDF (opens in new tab)
(with Parvin Lab)
 (opens in new tab)
(opens in new tab) Hill, McFaline-Figueroa et al. On the design of CRISPR-based single-cell molecular screens.
Nature Methods (2018)
PMID: 29457792 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
(opens in new tab) Matreyek, Starita et al. Multiplex assessment of protein variant abundance by massively parallel sequencing.
Nature Genetics (2018)
PMID: 29785012 (opens in new tab) | PDF (opens in new tab)
(with Fowler Lab)
2017
 (opens in new tab)
(opens in new tab) Gasperini, Findlay et al. CRISPR/Cas9-Mediated Scanning for Regulatory Elements Required for HPRT1 Expression via Thousands of Large, Programmed Genomic Deletions.
AJHG (2017)
 (opens in new tab)
(opens in new tab) Inoue, Kircher et al. A systematic comparison reveals substantial differences in chromosomal versus episomal encoding of enhancer activity.
Genome Research (2017)
PMID: 27831498 (opens in new tab) | PDF (opens in new tab)
(with Ahituv Lab)
2016
 (opens in new tab)
(opens in new tab) Gasperini, Starita, Shendure. The power of multiplexed functional analysis of genetic variants.
Nature Protocols (2016)
2015
 (opens in new tab)
(opens in new tab) Starita, Shendure, Fields et al. Massively Parallel Functional Analysis of BRCA1 RING Domain Variants.
Genetics (2015)
PMID: 25823446 (opens in new tab) | PDF (opens in new tab)
(with Fields Lab)
 (opens in new tab)
(opens in new tab) Kitzman, Starita et al. Massively parallel single-amino-acid mutagenesis.
Nature Methods (2015)
PMID: 25559584 (opens in new tab) | PDF (opens in new tab)
(with Fields Lab)
2014
 (opens in new tab)
(opens in new tab) Findlay, Boyle et al. Saturation editing of genomic regions by multiplex homology-directed repair.
Nature (2014)
2013
 (opens in new tab)
(opens in new tab) Smith, Taher L et al. Massively parallel decoding of mammalian regulatory sequences supports a flexible organizational model.
Nature Genetics (2013)
PMID: 23892608 (opens in new tab) | PDF (opens in new tab)
(with Ahituv, Ovcharenko Labs)
2012
 (opens in new tab)
(opens in new tab) Patwardhan, Hiatt et al. Massively parallel functional dissection of mammalian enhancers in vivo.
Nature Biotechnology (2012)
PMID: 22371081 (opens in new tab) | PDF (opens in new tab)
(with Pennacchio, Ahituv Labs)
2009
 (opens in new tab)
(opens in new tab) Patwardhan et al. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis.
Nature Biotechnology (2009)
