Genomic Approaches to Developmental Biology
(selected publications)
2026
 (opens in new tab)
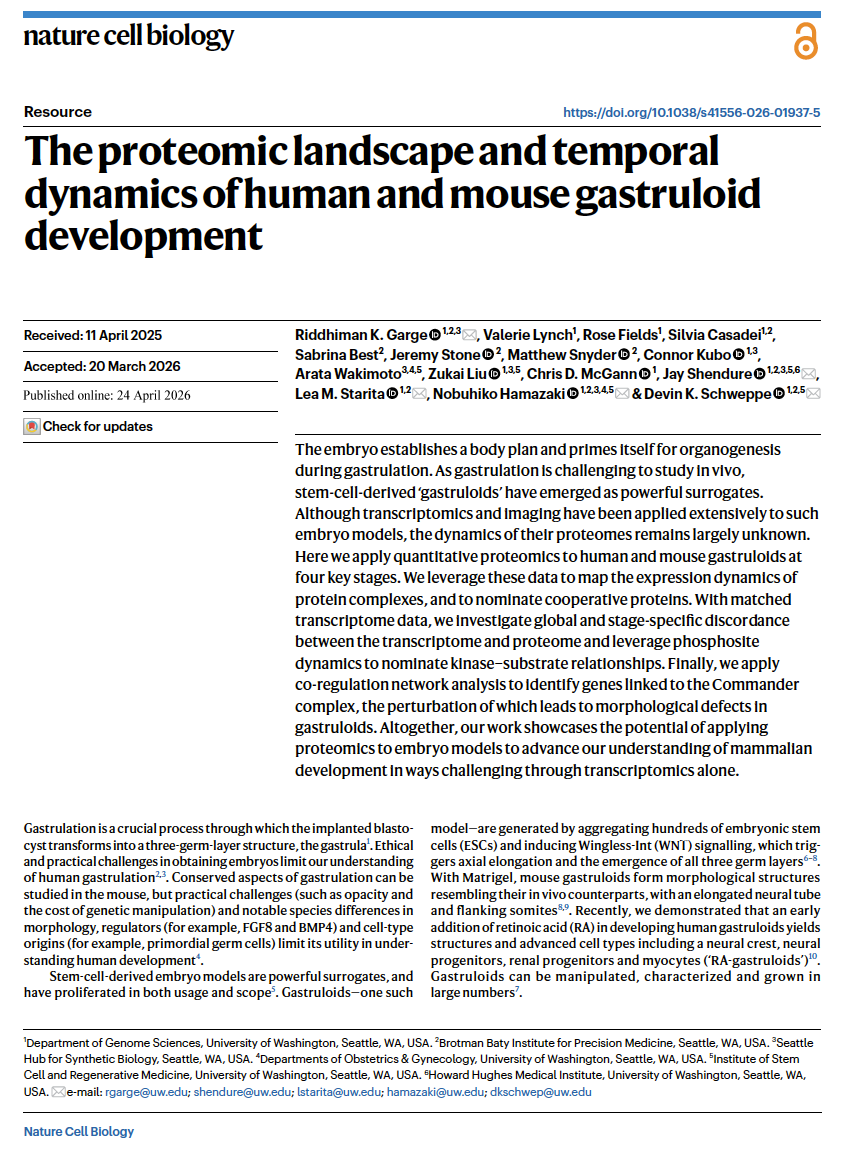
(opens in new tab) Garge et al. The proteomic landscape and temporal dynamics of human and mouse gastruloid development.
Nature Cell Biology (2026)
2024
 (opens in new tab)
(opens in new tab) Hamazaki, Yang et al. Retinoic acid induces human gastruloids with posterior embryo-like structures.
Nature Cell Biology (2024)
 (opens in new tab)
(opens in new tab) Chen, Choi et al. Symbolic recording of signalling and cis-regulatory element activity to DNA.
Nature (2024)
 (opens in new tab)
(opens in new tab) Qiu, Martin, Welsh et al. A single-cell time-lapse of mouse prenatal development from gastrula to birth.
Nature (2024)
2023
 (opens in new tab)
(opens in new tab) Domcke, Shendure. A reference cell tree will serve science better than a reference cell atlas.
Cell (2023)
 (opens in new tab)
(opens in new tab) Huang, Henck, Qiu et al. Single-cell, whole embryo phenotyping of mammalian developmental disorders.
Nature (2023)
PMID: 37968388 (opens in new tab) | PDF (opens in new tab)
(with Spielmann and Cao Labs)
 (opens in new tab)
(opens in new tab) Chiou, Huang et al. A single-cell multi-omic atlas spanning the adult rhesus macaque brain.
Science Advances (2023)
2022
 (opens in new tab)
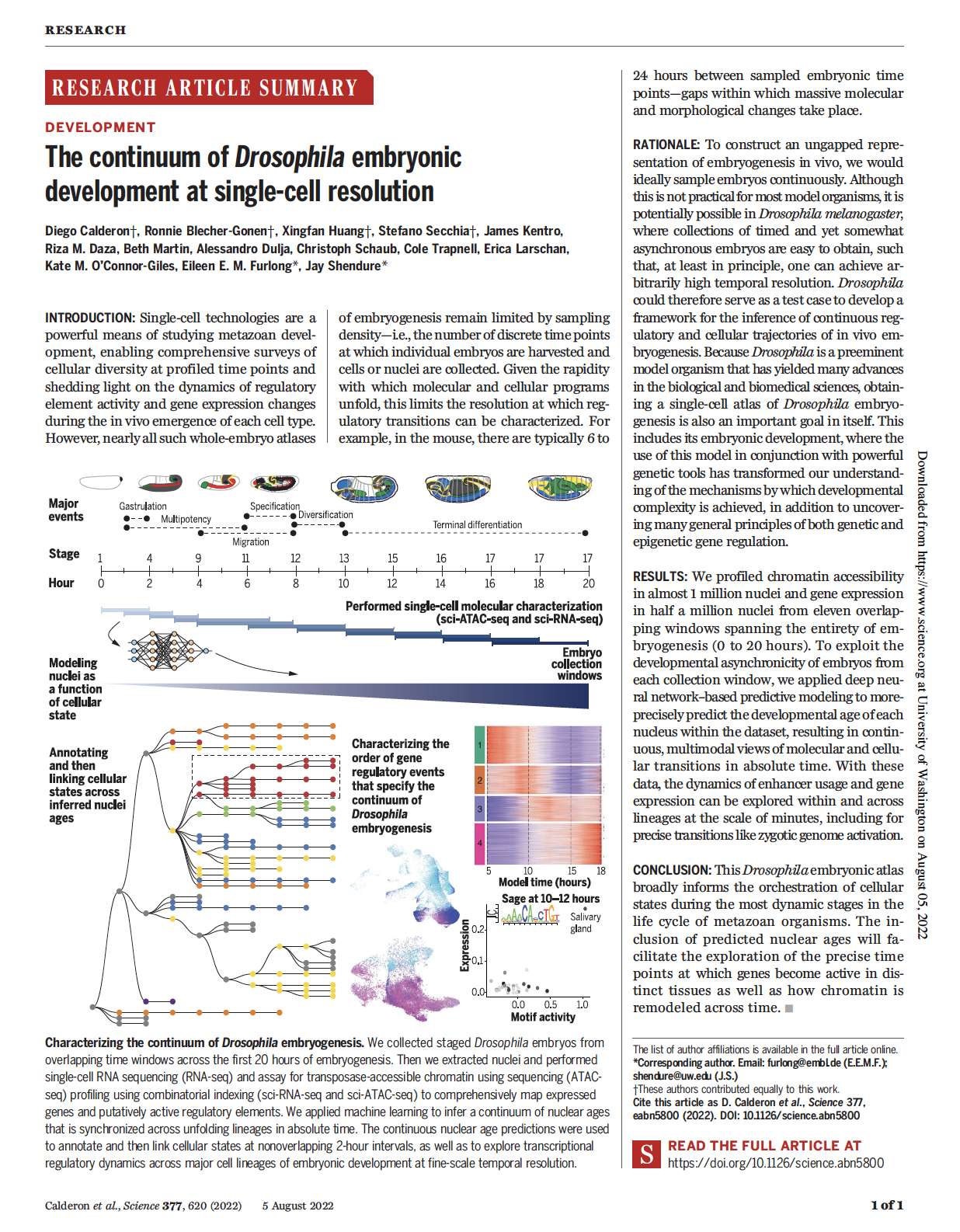
(opens in new tab) Calderon, Huang et al. The continuum of Drosophila embryonic development at single-cell resolution.
Science (2022)
PMID: 35926038 (opens in new tab) | PDF (opens in new tab)
(with Furlong Lab)
 (opens in new tab)
(opens in new tab) Choi et al. A time-resolved, multi-symbol molecular recorder via sequential genome editing.
Nature (2022)
 (opens in new tab)
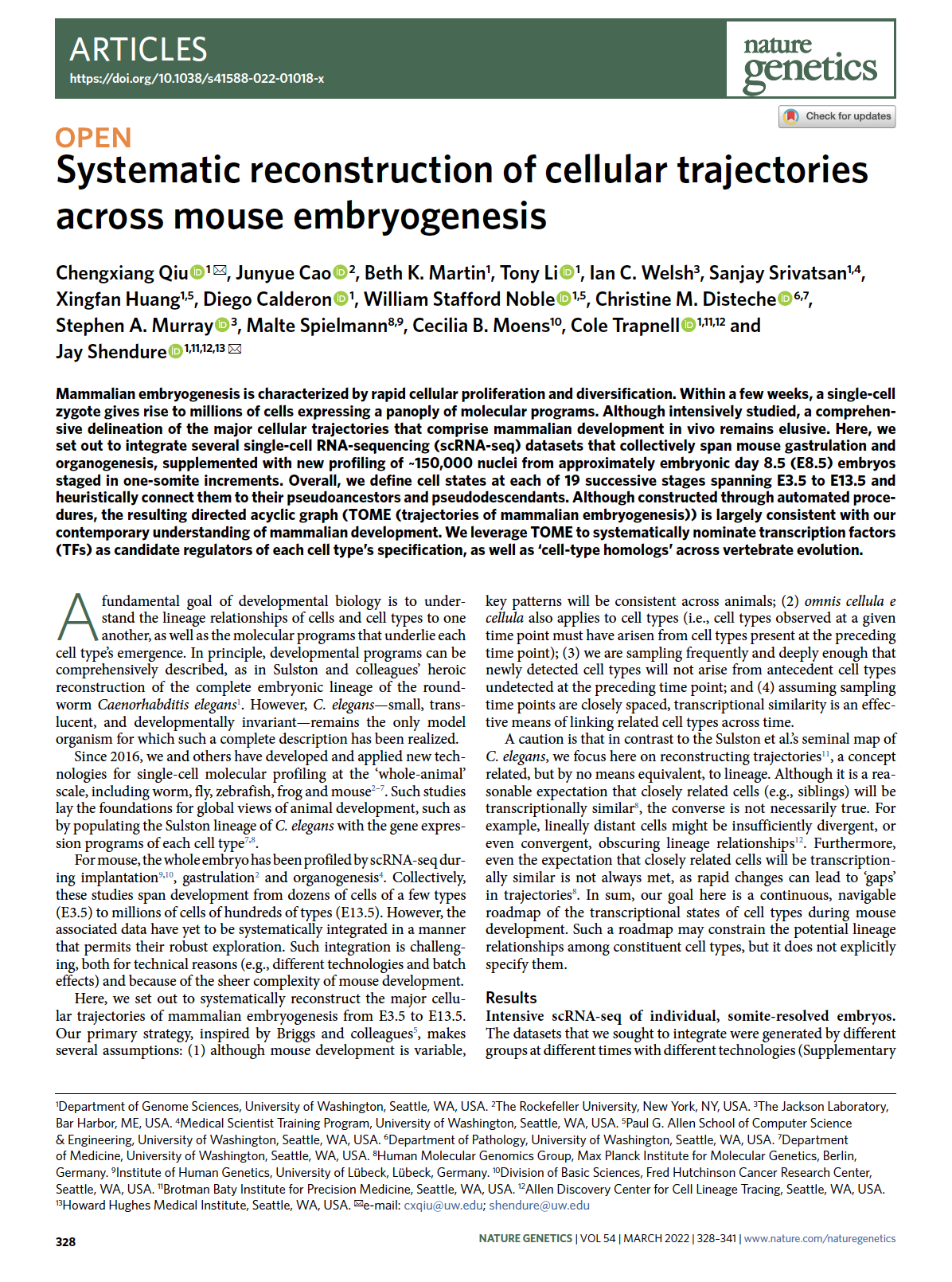
(opens in new tab) Qiu et al. Systematic reconstruction of cellular trajectories across mouse embryogenesis.
Nature Genetics (2022)
2021
.png) (opens in new tab)
(opens in new tab) Srivatsan, Regier et al. Embryo-scale, single-cell spatial transcriptomics.
Science (2021)
PMID: 34210887 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
(opens in new tab) Agarwal, Darwin-Lopez et al. The landscape of alternative polyadenylation in single cells of the developing mouse embryo.
Nature Communications (2021)
2020
 (opens in new tab)
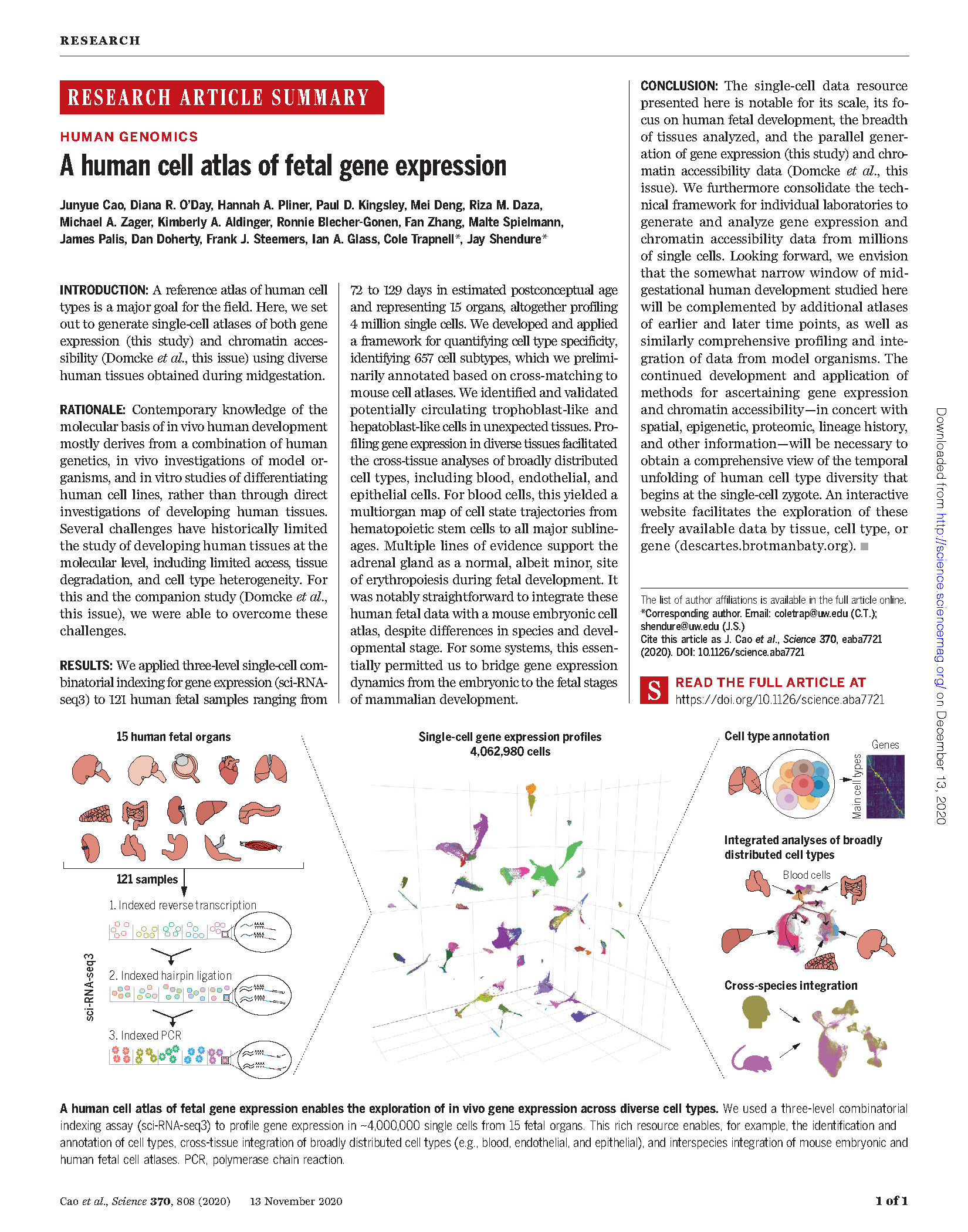
(opens in new tab) Cao et al. A human cell atlas of fetal gene expression.
Science (2020)
PMID: 33184181 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
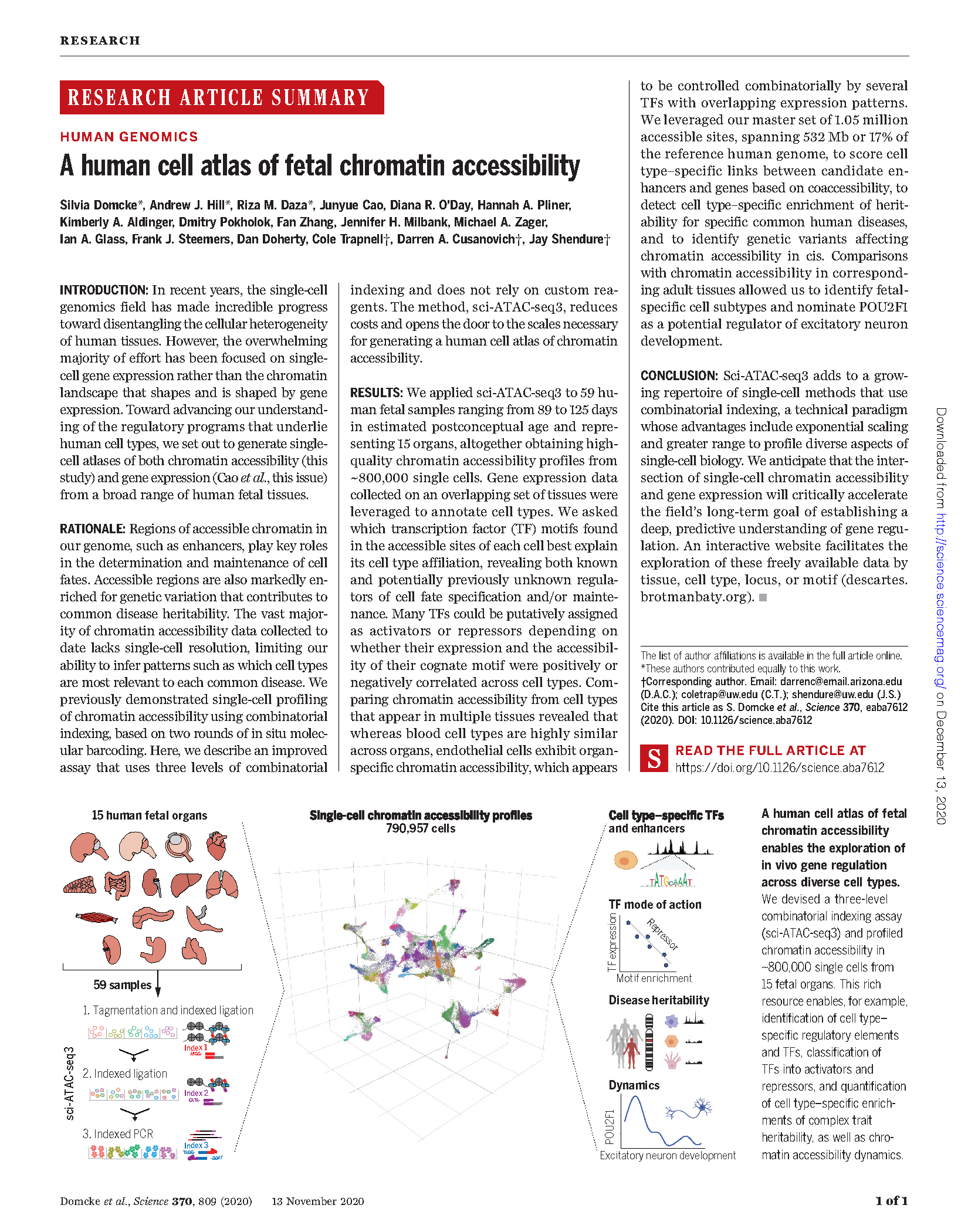
(opens in new tab) Domcke, Hill, Daza et al. A human cell atlas of fetal chromatin accessibility.
Science (2020)
PMID: 33184180 (opens in new tab) | PDF (opens in new tab)
(with Cusanovich and Trapnell Labs)
 (opens in new tab)
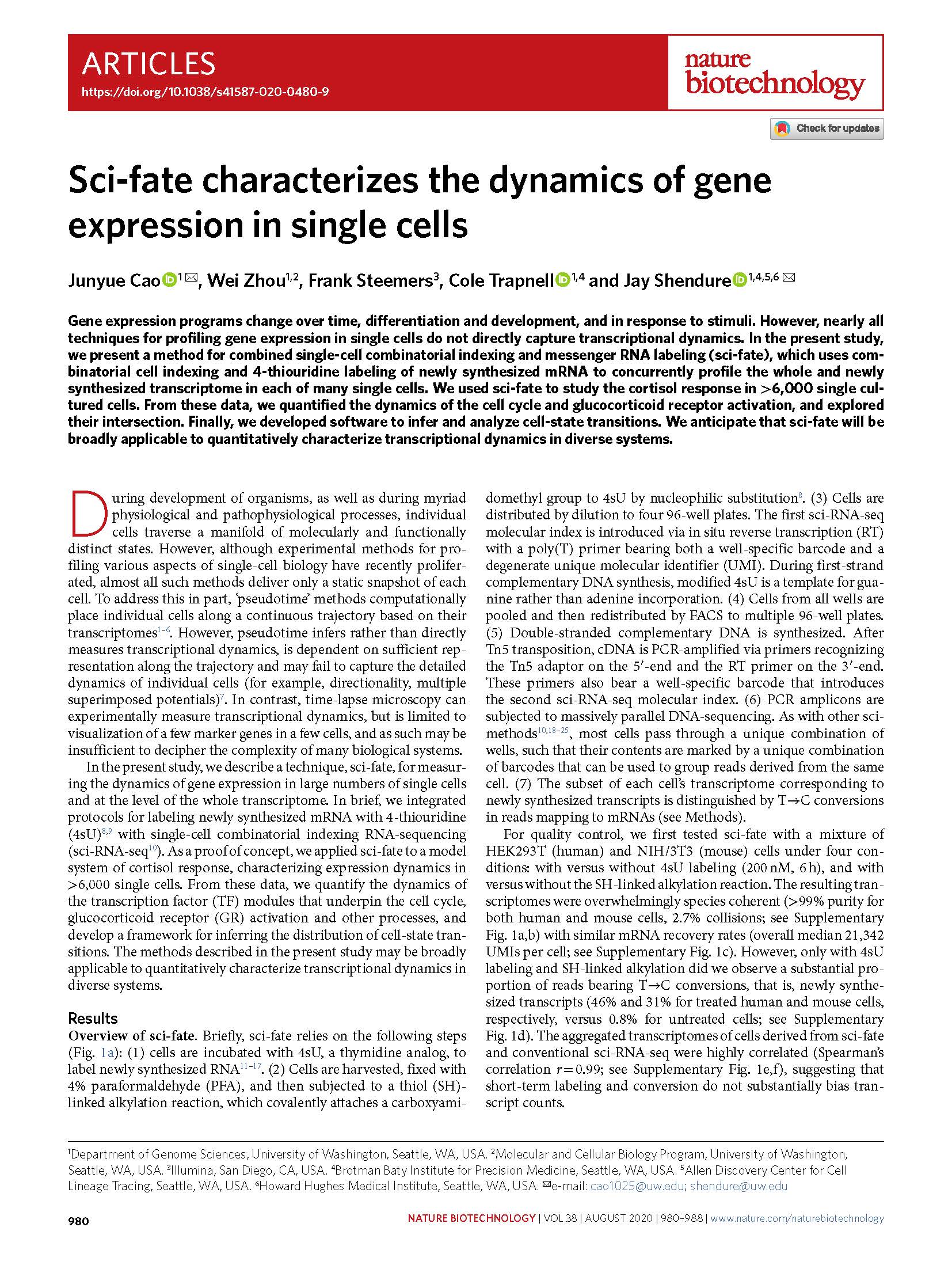
(opens in new tab) Cao et al. Sci-fate characterizes the dynamics of gene expression in single cells.
Nature Biotechnology (2020)
2019
 (opens in new tab)
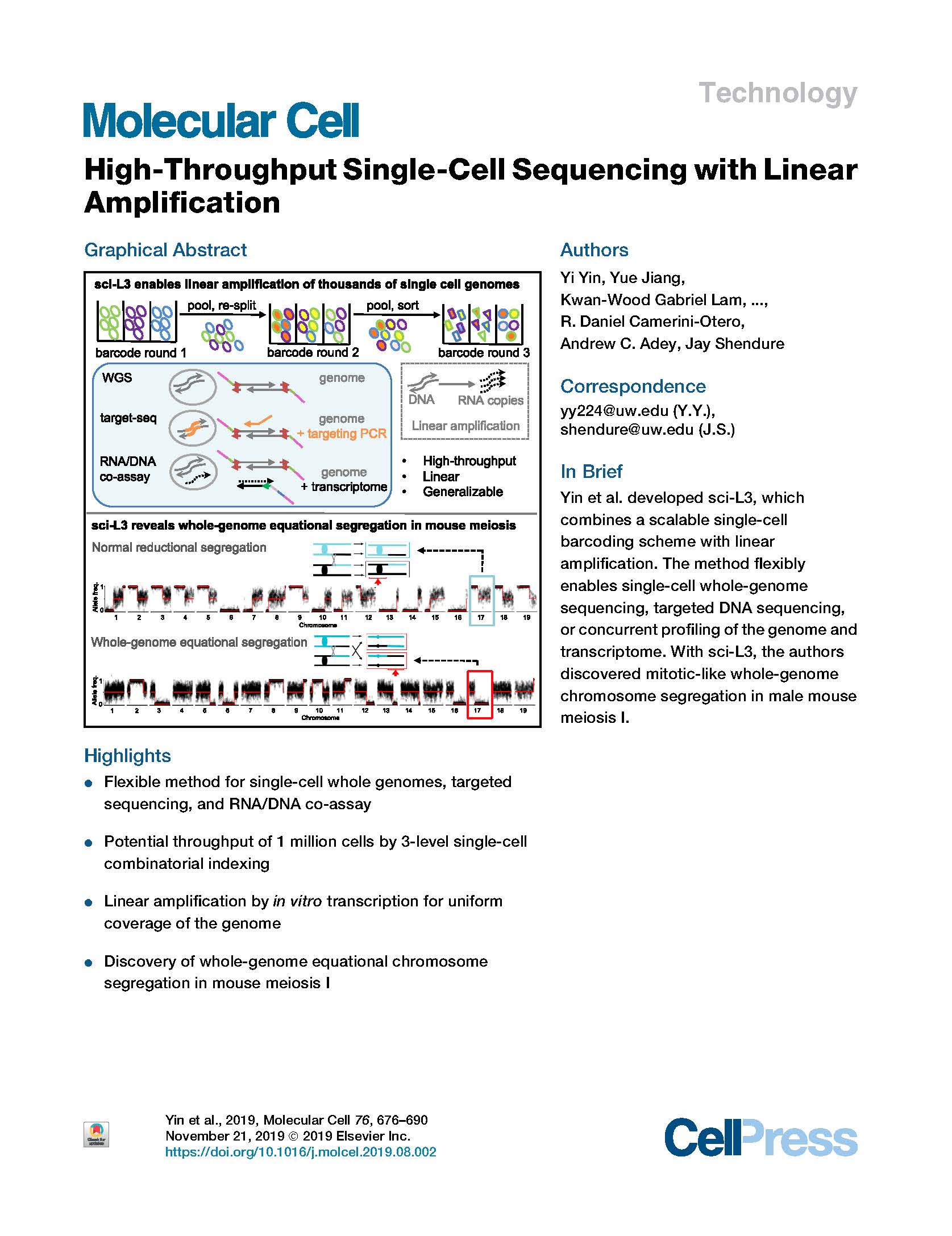
(opens in new tab) Yin et al. High-Throughput Single-Cell Sequencing with Linear Amplification.
Molecular Cell (2019)
 (opens in new tab)
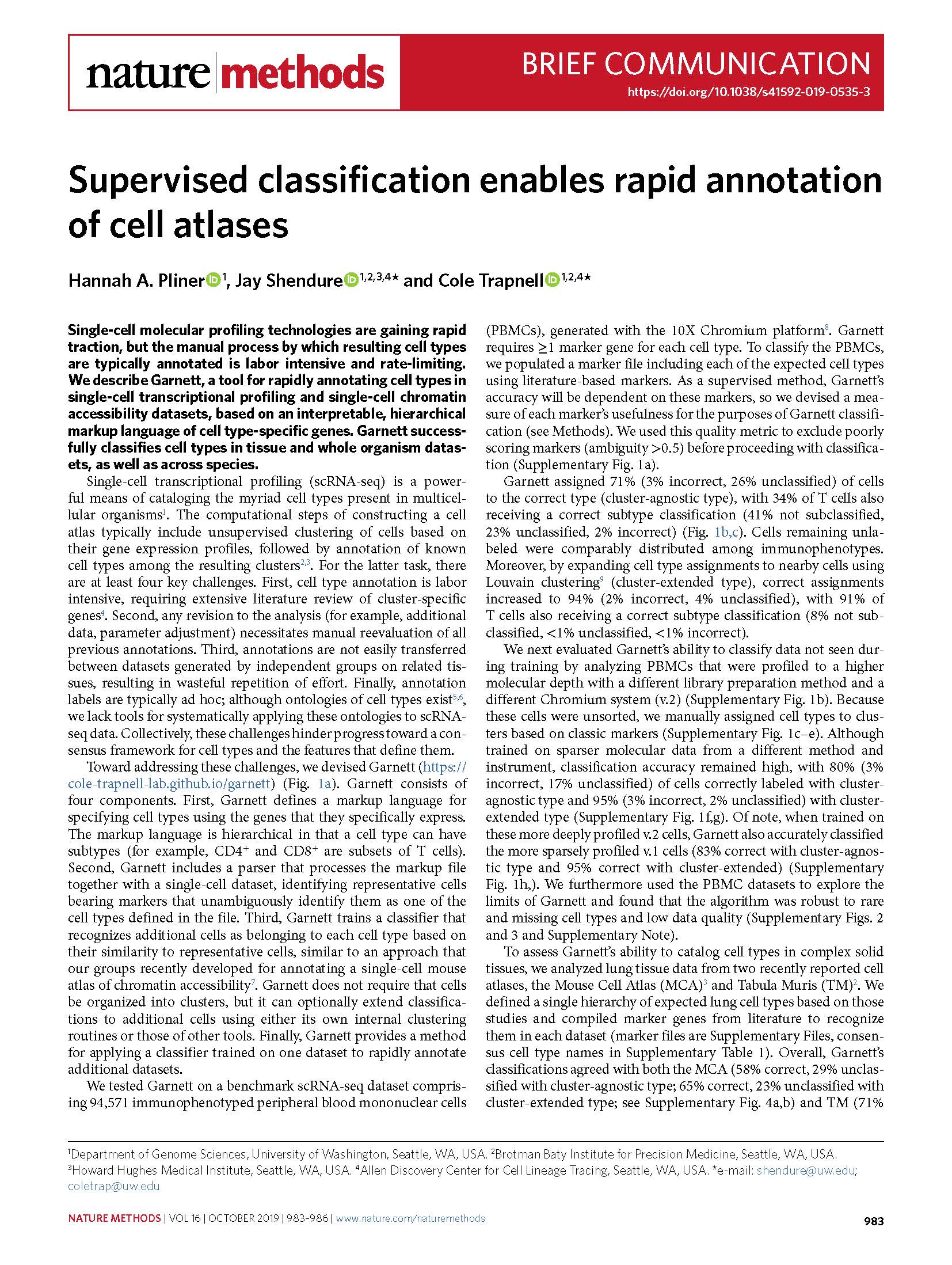
(opens in new tab) Pliner et al. Supervised classification enables rapid annotation of cell atlases.
Nature Methods (2019)
PMID: 31501545 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
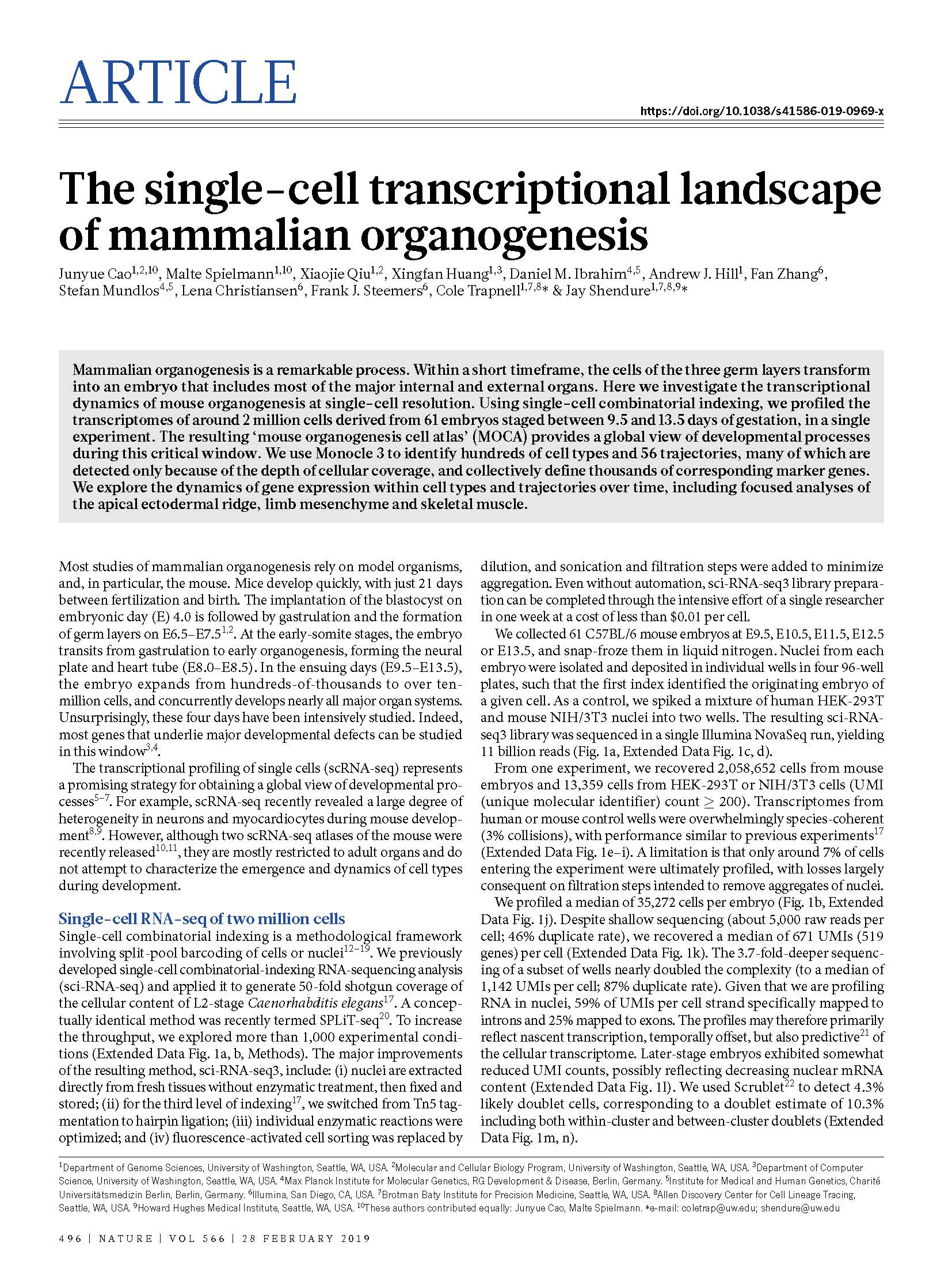
(opens in new tab) Cao, Spielmann et al. The single-cell transcriptional landscape of mammalian organogenesis.
Nature (2019)
PMID: 30787437 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
2018
 (opens in new tab)
(opens in new tab) Cao et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells.
Science (2018)
PMID: 30166440 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
(opens in new tab) Cusanovich, Hill et al. A Single-Cell Atlas of In Vivo Mammalian Chromatin Accessibility.
Cell (2018)
PMID: 30078704 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
(opens in new tab) Pliner et al. Cicero Predicts cis-Regulatory DNA Interactions from Single-Cell Chromatin Accessibility Data.
Molecular Cell (2018)
PMID: 30078726 (opens in new tab) | PDF (opens in new tab)
(with Trapnell Lab)
 (opens in new tab)
(opens in new tab) Cusanovich, Reddington, Garfield et al. The cis-regulatory dynamics of embryonic development at single-cell resolution.
Nature (2018)
PMID: 29539636 (opens in new tab) | PDF (opens in new tab)
(with Furlong Lab)
2017
 (opens in new tab)
(opens in new tab) Cao, Packer et al. Comprehensive single cell transcriptional profiling of a multicellular organism by combinatorial indexing.
Science (2017)
PMID: 28818938 (opens in new tab) | PDF (opens in new tab)
(with Trapnell and Waterston Labs)
2016
 (opens in new tab)
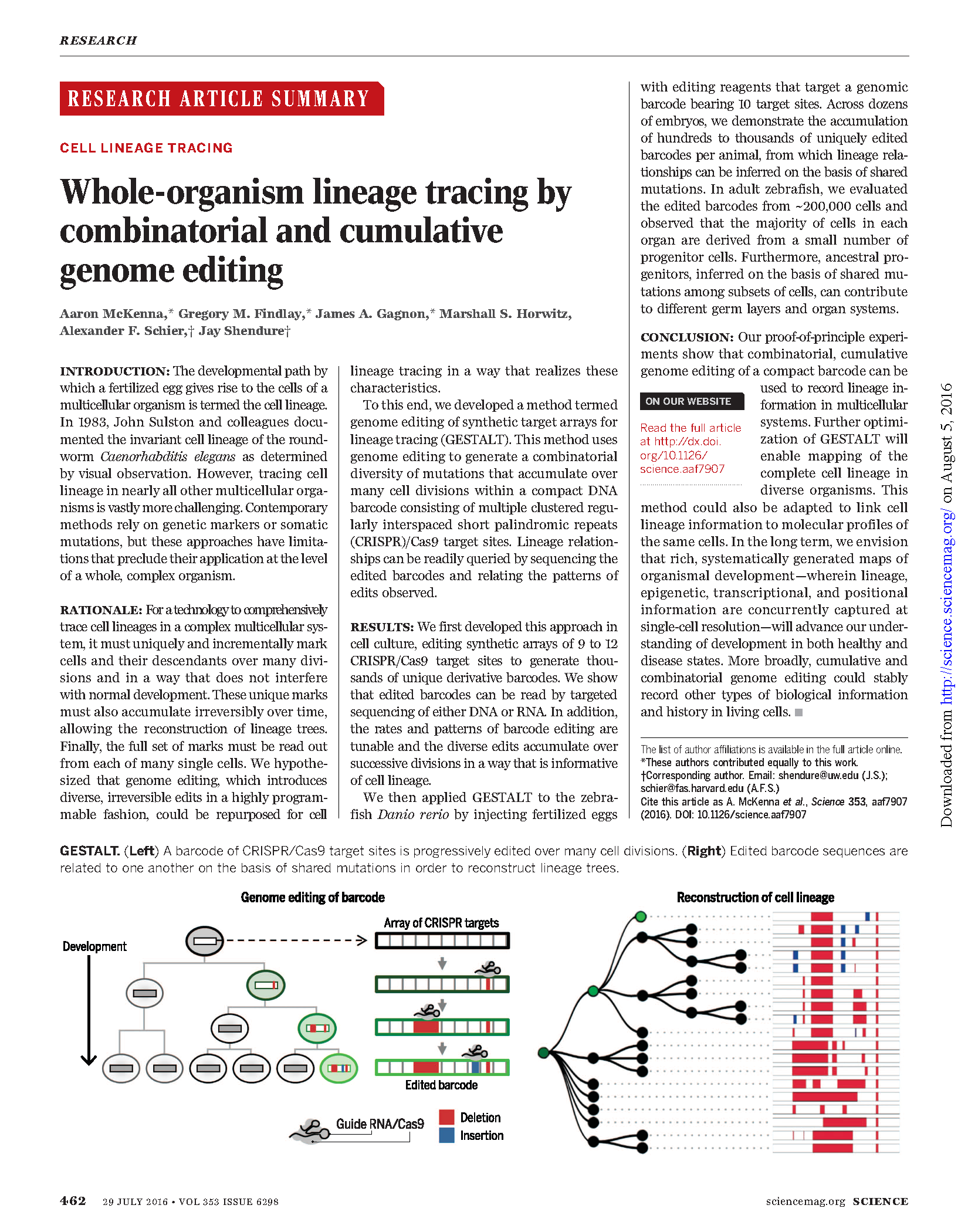
(opens in new tab) McKenna, Findlay, Gagnon et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing.
Science (2016)
PMID: 27229144 (opens in new tab) | PDF (opens in new tab)
(with Schier Lab)