Software
Shendure Lab GitHub (opens in new tab)
Public GitHub repositories for assorted lab projects.
Other Software
 (opens in new tab)
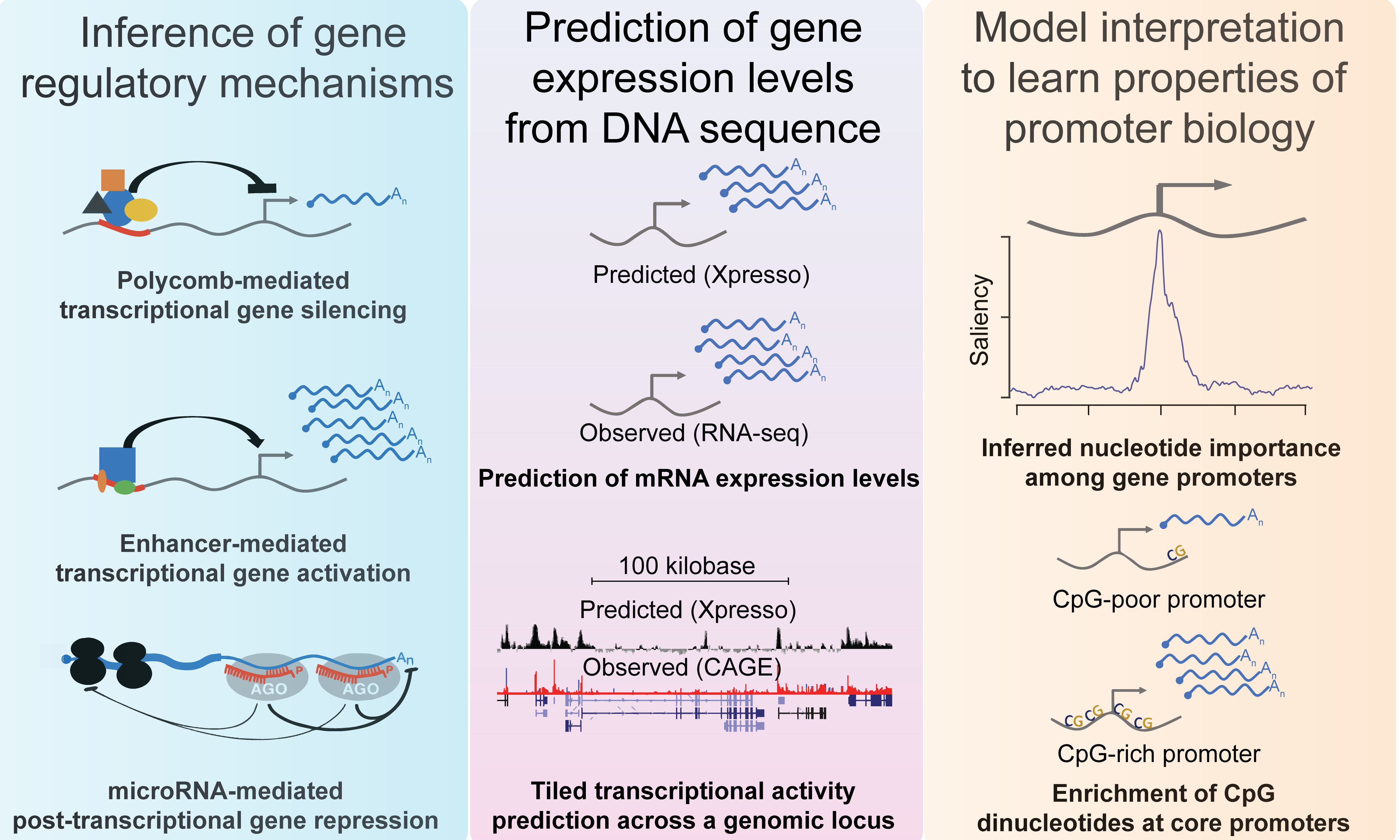
(opens in new tab) Xpresso (opens in new tab)
Predicting gene expression levels from DNA sequences
Xpresso is a software suite whose goal is to predict gene expression levels and transcriptional activity from genomic sequences. It is trained using convolutional neural networks. Pre-trained models are available for the human, mouse, and several cell types for these species.
 (opens in new tab)
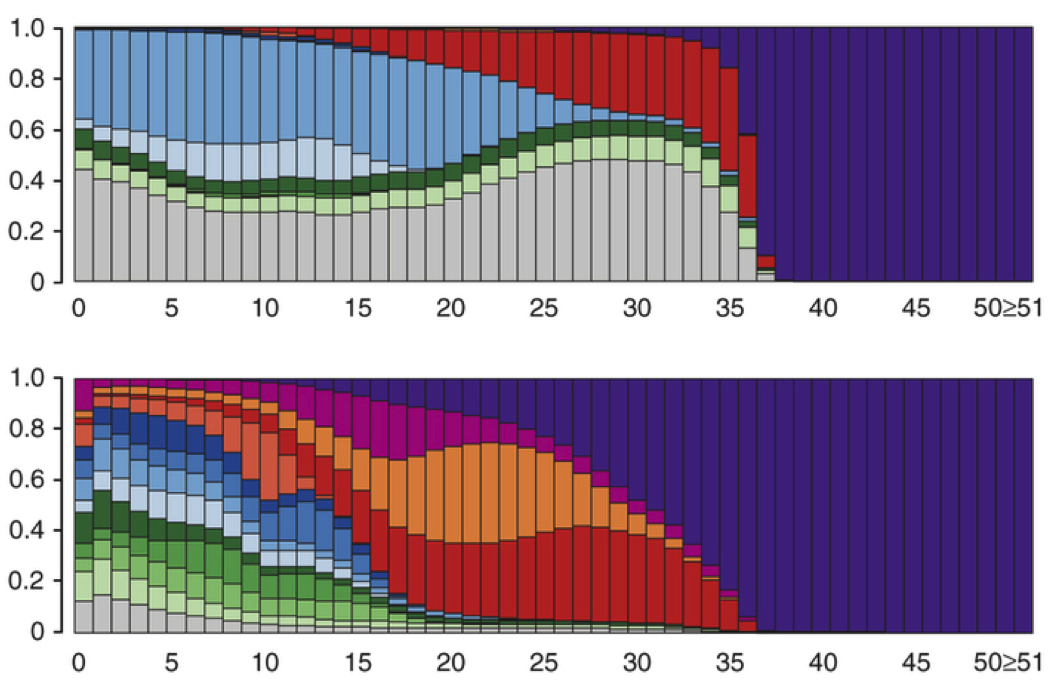
(opens in new tab) GESTALT (opens in new tab)
Genome Editing of Synthetic Target Arrays for Lineage Tracing
GESTALT is a method that uses genome editing to progressively introduce and accumulate diverse mutations in a DNA barcode over multiple rounds of cell division, thereby recording cell lineage relationships in the patterns of mutations shared between cells.
 (opens in new tab)
(opens in new tab) CADD (opens in new tab)
Combined Annotation Dependent Depletion
CADD is method that objectively weights and integrates diverse genomic annotations to a single, phred-scaled metric. Pre-computed CADD-based scores (C-scores) for all 8.6 billion possible single nucleotide variants (SNVs) of the human reference genome, and a tool for scoring of short insertions/deletions are available.
 (opens in new tab)
(opens in new tab) MIPgen (opens in new tab)
Optimized Modeling and Design of Molecular Inversion Probes for Targeted Resequencing
Molecular inversion probes (MIPs) enable cost-effective multiplex targeted gene resequencing in large cohorts. MIPgen is a user-friendly package that simplifies the process of designing custom MIP assays to arbitrary targets. New logistic and SVM-derived models enable in silico predictions of assay success.
 (opens in new tab)
(opens in new tab) LACHESIS (opens in new tab)
Genome Assembly with Hi-C-based Contact Probability Maps
LACHESIS is method that exploits contact probability map data (e.g. from Hi-C) for chromosome-scale de novo genome assembly.
 (opens in new tab)
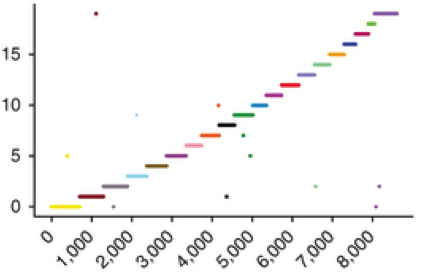
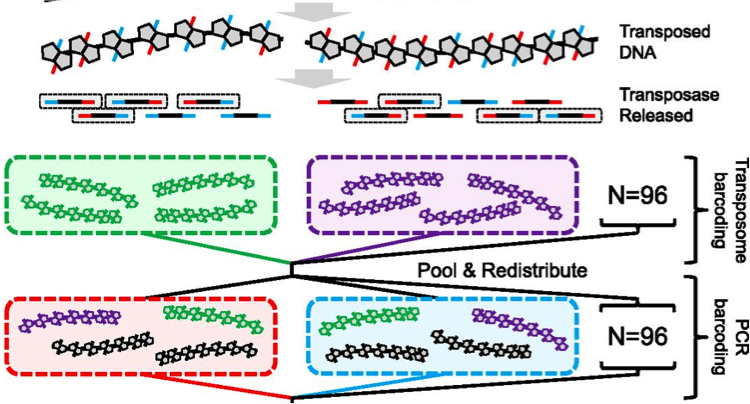
(opens in new tab) fragScaff (opens in new tab)
Genome Assembly with Contiguity Preserving Transposition
Contiguity preserving transposition and sequencing (CPT-seq) is an entirely in vitro means of generating libraries comprised of 9216 indexed pools. FragScaff leverages coincidences between the content of different pools as a source of contiguity information for scaffolding de novo genome assemblies.
 (opens in new tab)
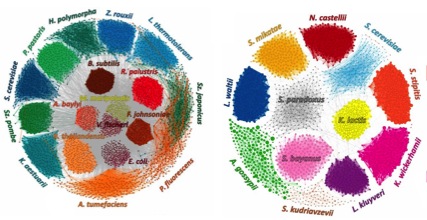
(opens in new tab) MetaPhase (opens in new tab)
Species-level Deconvolution of Metagenome Assemblies with Hi-C-based Contact Probability Maps
MetaPhase is software that exploits chromatin-level contact probability maps, e.g., as generated by the Hi-C method, to reconstruct the individual genomes of microbial species present within a mixed sample.